Salt (1.5 M) avoidance test in DAN-larvae under red and blue light (N=10)
on Sunday, April 12th, 2026 2:07 | by Christoph Kumpfmüller
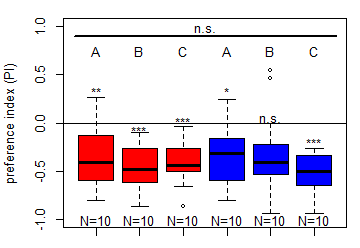
Category: DAN, genetics, Larvae, Mushroom Body, Optogenetics | No Comments
T4/T5 lines test crossings
on Monday, March 30th, 2026 12:18 | by Radostina Lyutova
Different T4/T5 driver lines are crossed with different effector lines to assess the viability of the F1 progeny
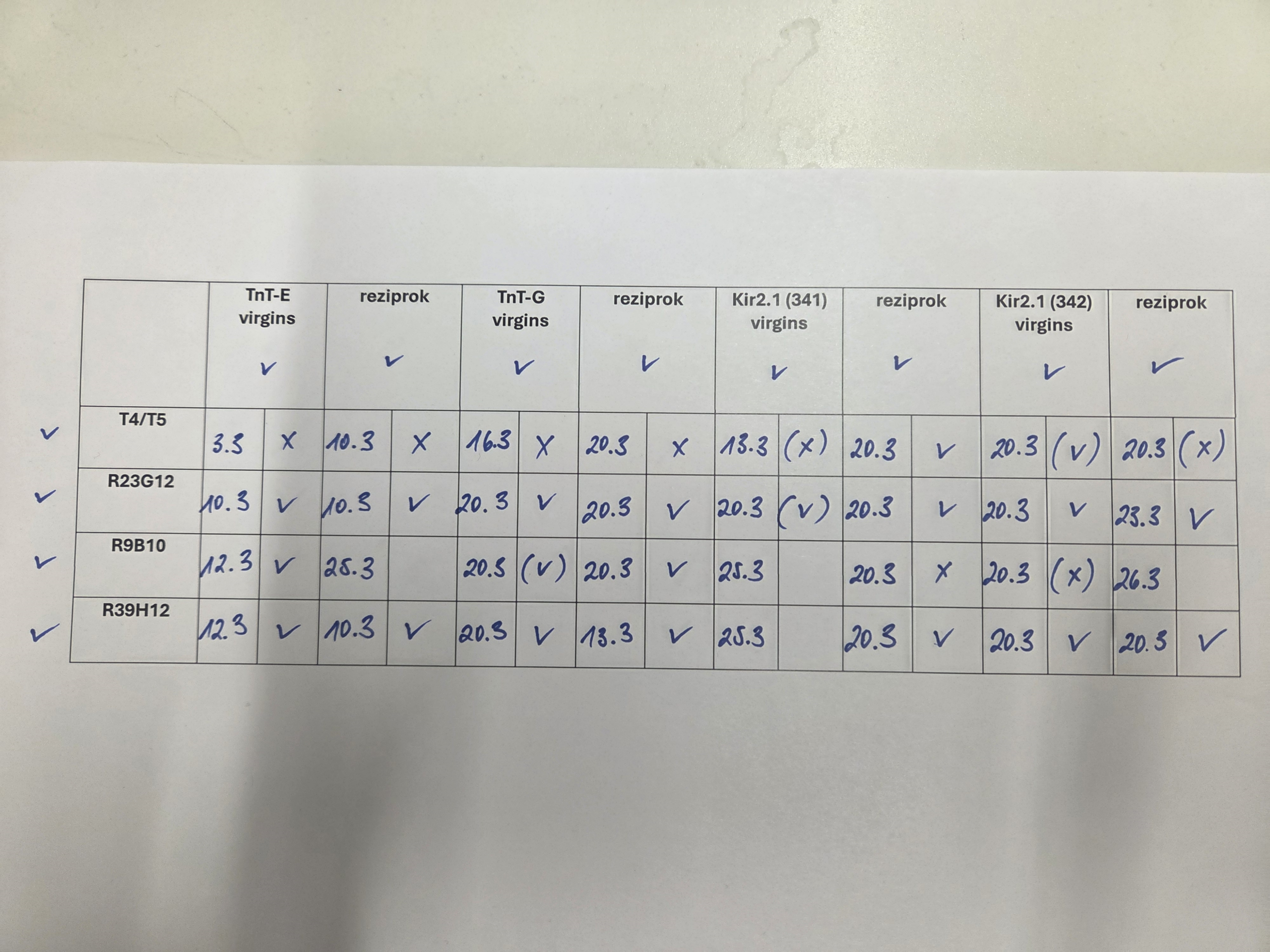
Category: crosses, T4/T5 | No Comments
Finished salt (1.5 M) avoidance tests in larvae (Dop2R) under red and blue light
on Thursday, March 26th, 2026 2:25 | by Christoph Kumpfmüller
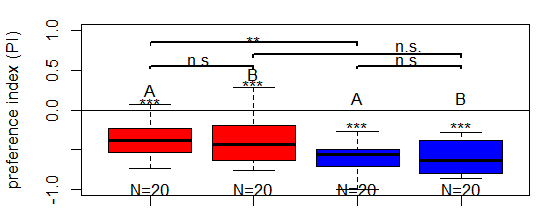
ARL <-> ABL: 0.009
BRL <-> BBL: 0.053
Category: Anatomy, crosses, genetics, Larvae, Mushroom Body, Optogenetics | No Comments
T4/T5 lines test crossings
on Monday, March 16th, 2026 12:04 | by Radostina Lyutova
Different T4/T5 driver lines are crossed with different effector lines to assess the viability of the F1 progeny
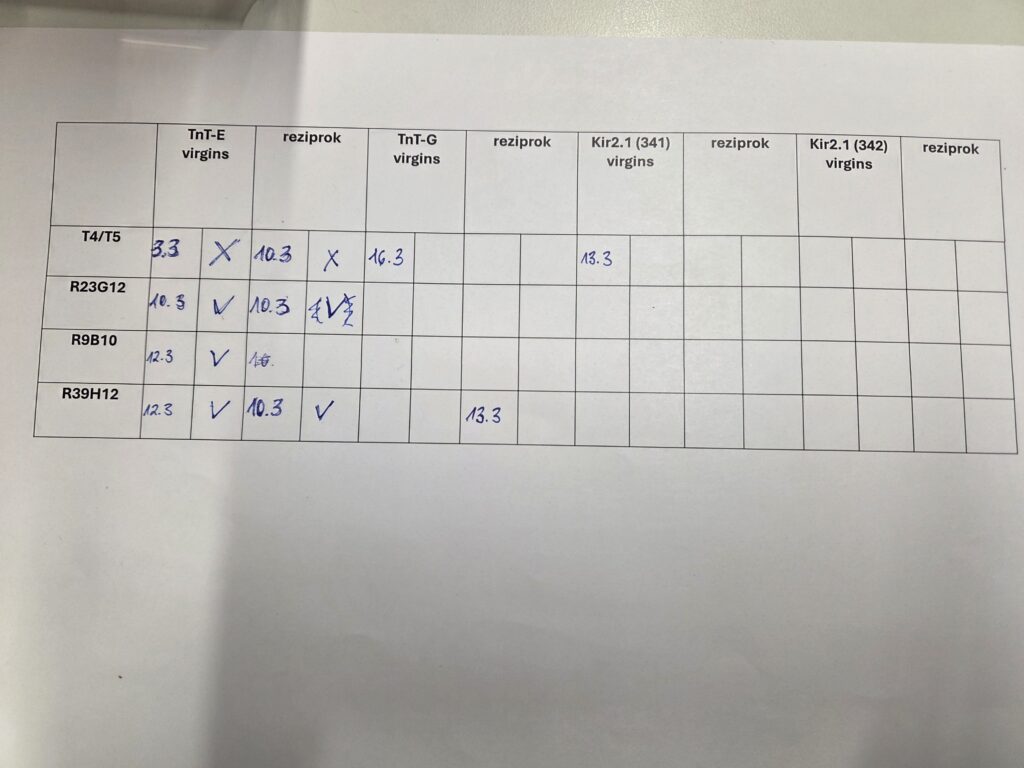
Category: crosses | No Comments
Salt (1.5 M) avoidance tests in larvae (Dop2R) under red and blue light
on Sunday, March 15th, 2026 4:49 | by Christoph Kumpfmüller
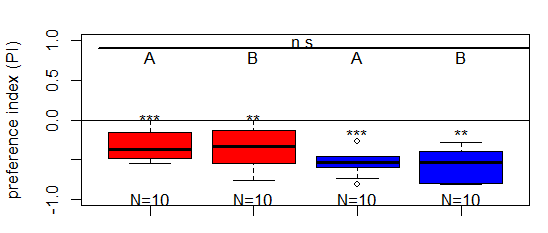
Category: crosses, genetics, Larvae, Optogenetics | No Comments
Salt (1.5 M) avoidance tests in larvae (DANc1) under blue and red light (N = 20)
on Sunday, March 8th, 2026 5:55 | by Christoph Kumpfmüller
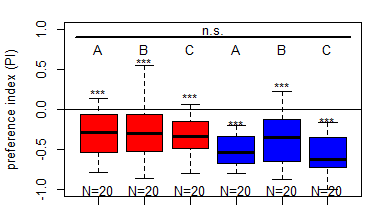
Category: crosses, genetics, Larvae, Mushroom Body, Optogenetics | No Comments
Salt (1.5 M) avoidance tests in larvae (DANc1) under blue and red light (N = 12)
on Sunday, March 1st, 2026 4:56 | by Christoph Kumpfmüller
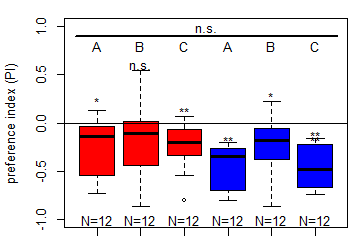
Category: crosses, genetics, Larvae, Mushroom Body, Optogenetics | No Comments
Finished salt (1.5 M) avoidance test in MB065b-[DAN_f1/DAN_c1] larvae under red and blue light
on Sunday, February 22nd, 2026 6:16 | by Christoph Kumpfmüller
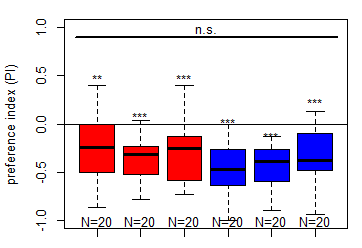
Category: crosses, DAN, Larvae, Mushroom Body, Optogenetics | No Comments
Salt (1.5 M) avoidance test in MB065b-[DAN_f1/DAN_c1] larvae under red and blue light
on Saturday, February 14th, 2026 3:15 | by Christoph Kumpfmüller
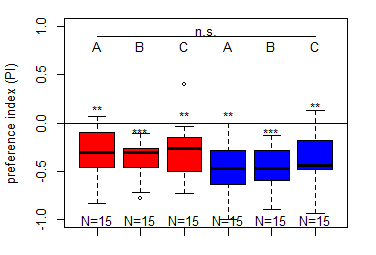
Category: crosses, DAN, Larvae, Mushroom Body, Optogenetics | No Comments
Confocal images Daniel’s samples
on Monday, February 9th, 2026 12:33 | by Fridrik Kjartansson
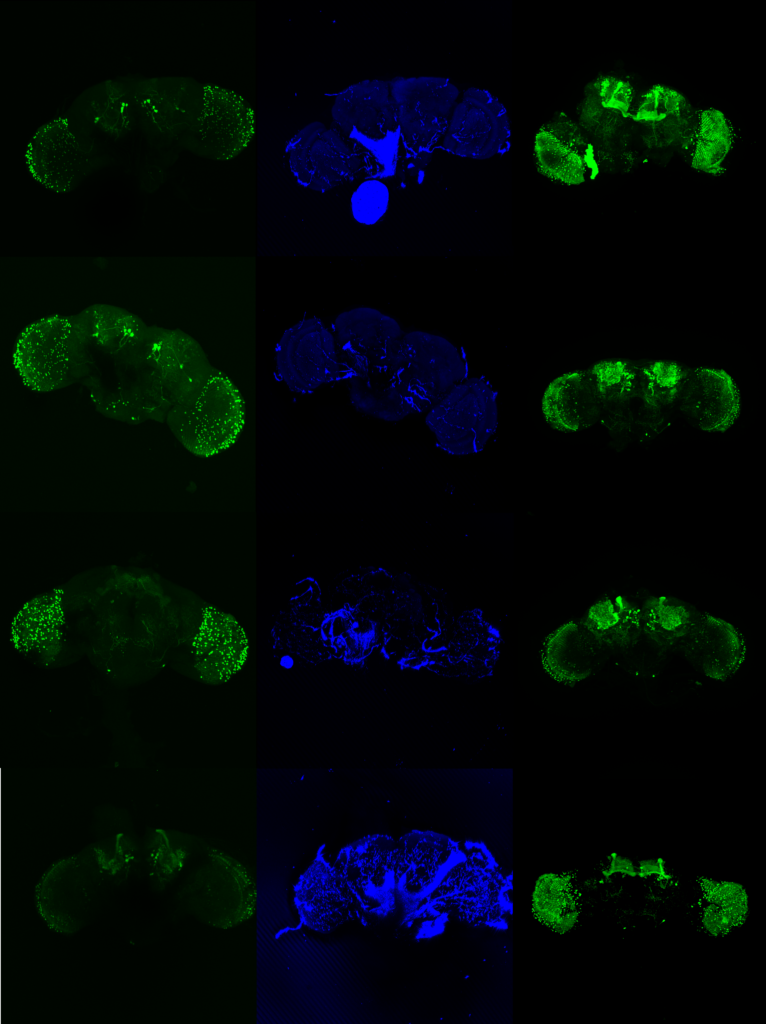
Category: crosses, Mushroom Body | No Comments
