Creating gRNAs via PCR
on Friday, July 7th, 2023 6:38 | by Isabel Stark
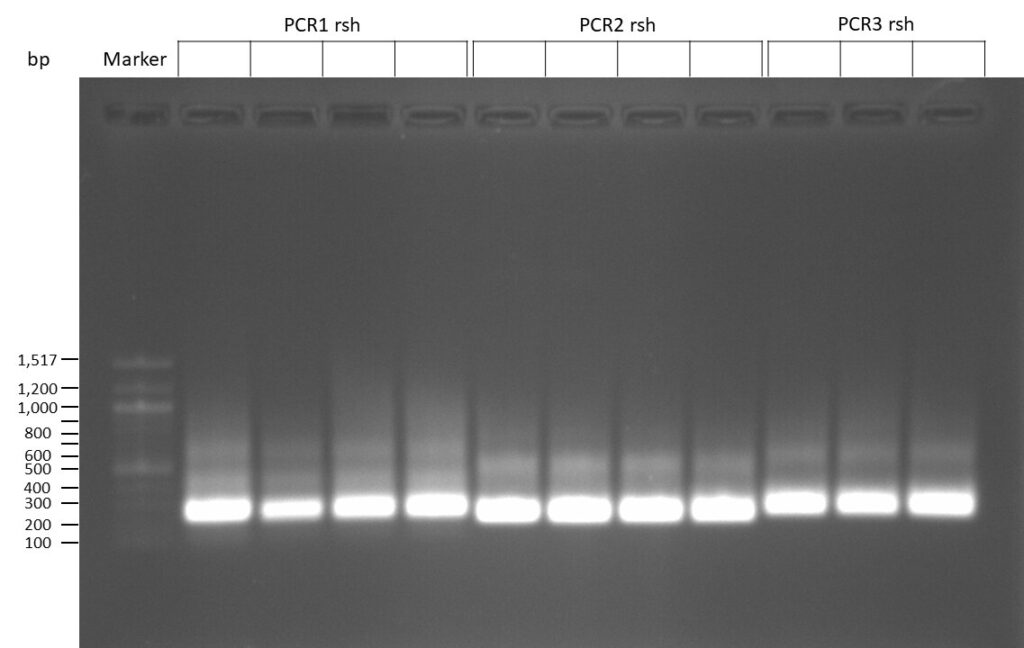
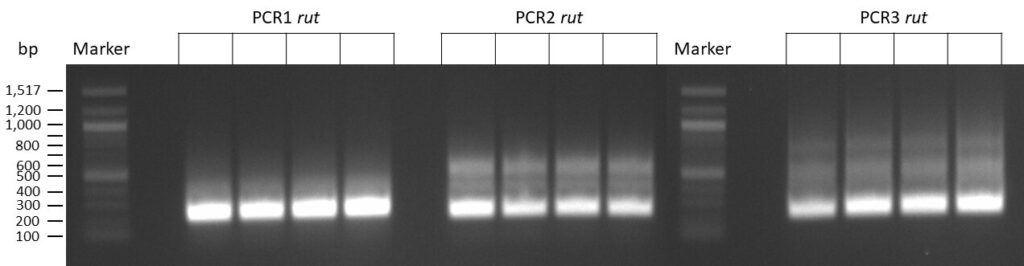
The template pCFD6 was used with a concentration of 640 pg/µl.
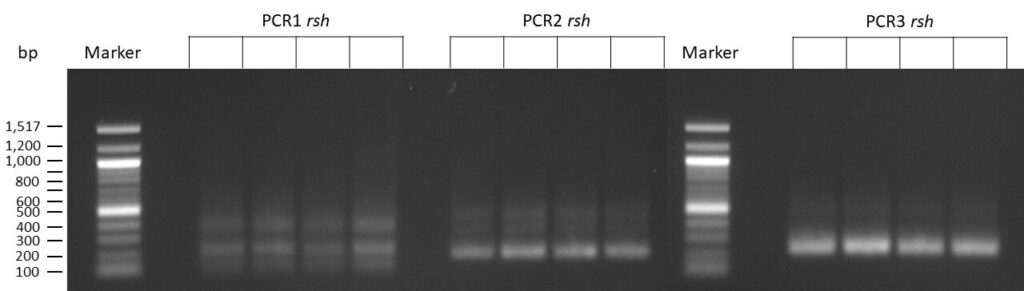
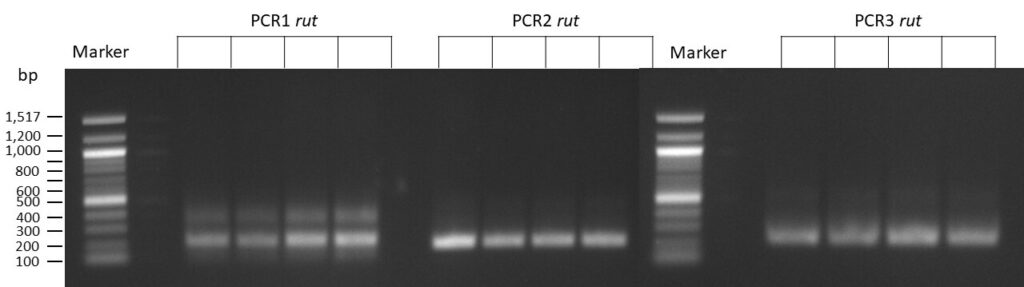
The template pCFD6 was used with a concentration of 64 pg/µl (1:10 dilution).
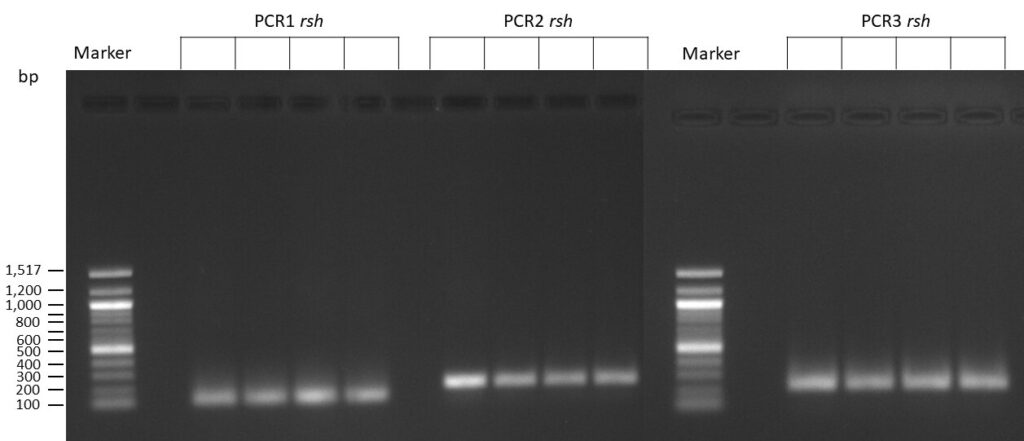
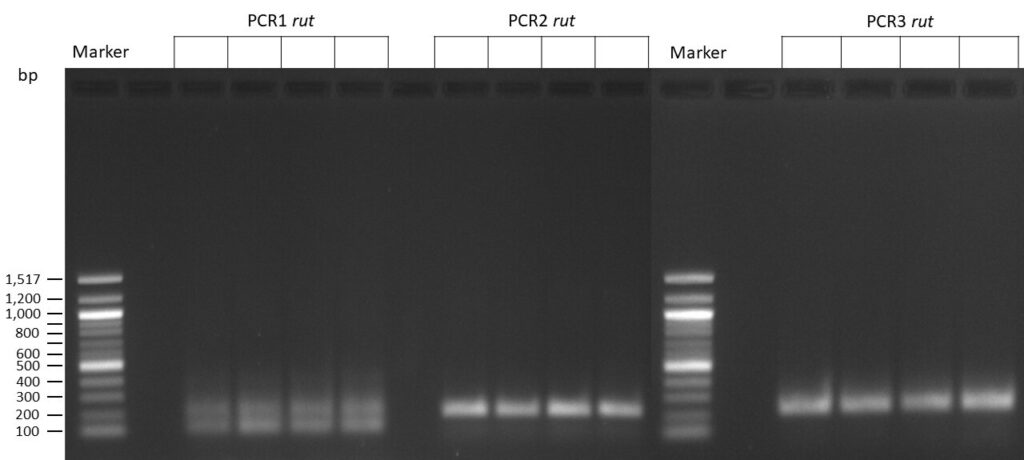
The template pCFD6 was used with a concentration of 64 pg/µl (1:10 dilution).
50µl of 5xQ5 High GC Enhancer was added to the PCR mix.
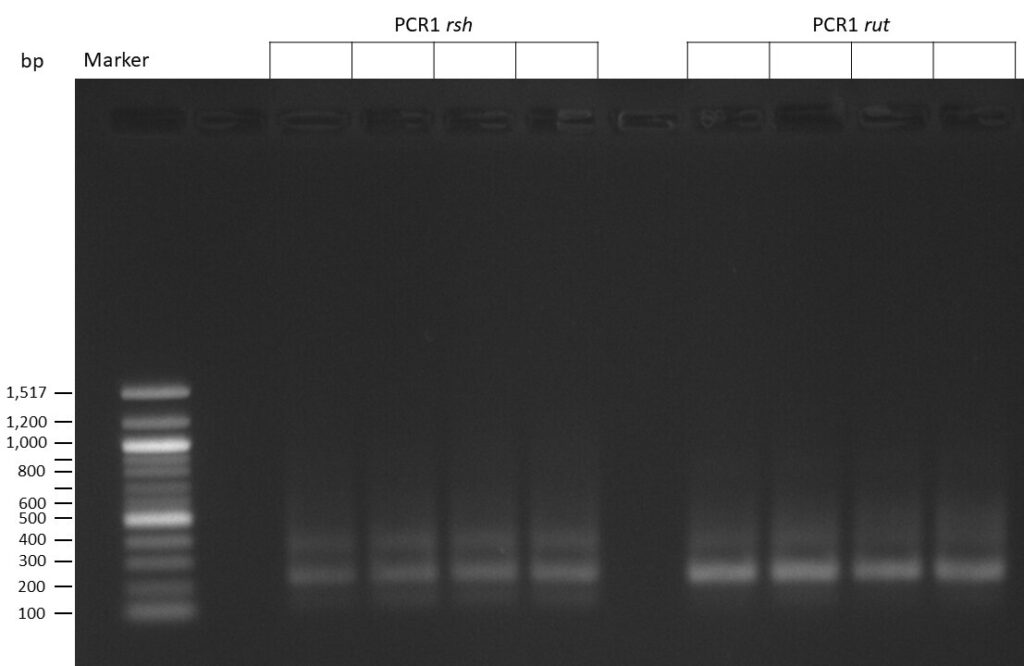
The template pCFD6 was used with a concentration of 128 pg/µl (1:5 dilution).
50µl of 5xQ5 High GC Enhancer was added to the PCR mix.
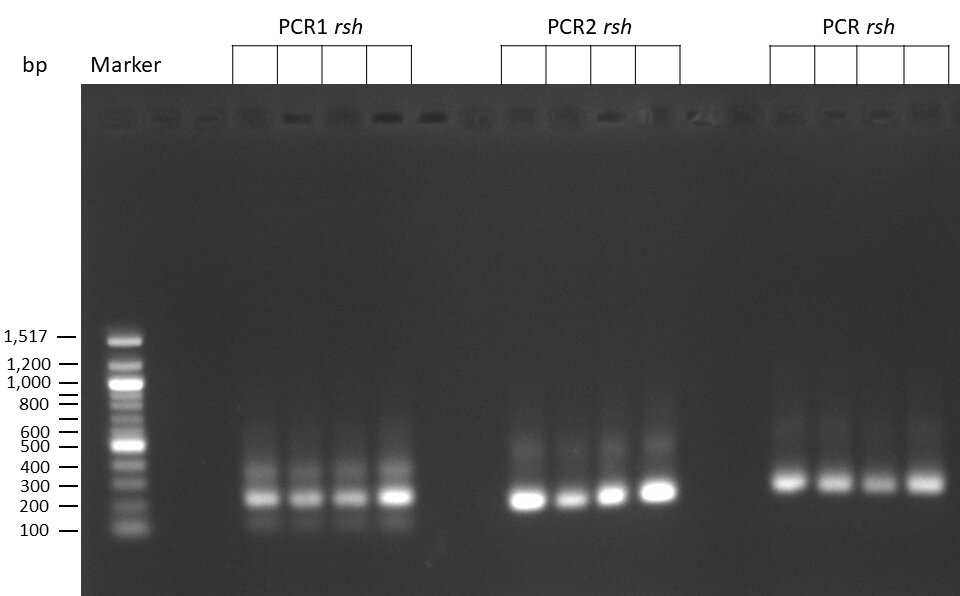
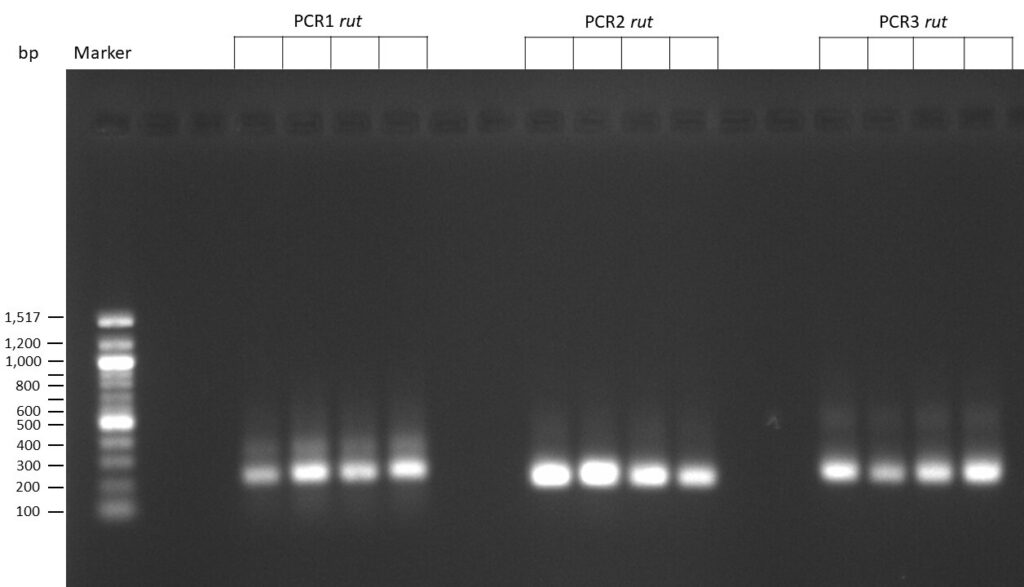
The template pCFD6 was used with a concentration of 640 pg/µl.
The Phusion DNA Polymerase was used instead of the Q5 High-Fidelity DNA Polymerase.
Category: genetics, Memory, Operant learning, operant self-learning, Radish, Uncategorized

Leave a Reply