Salt Avoidance CantonS
on Monday, April 14th, 2025 1:37 | by Eva Schächtl
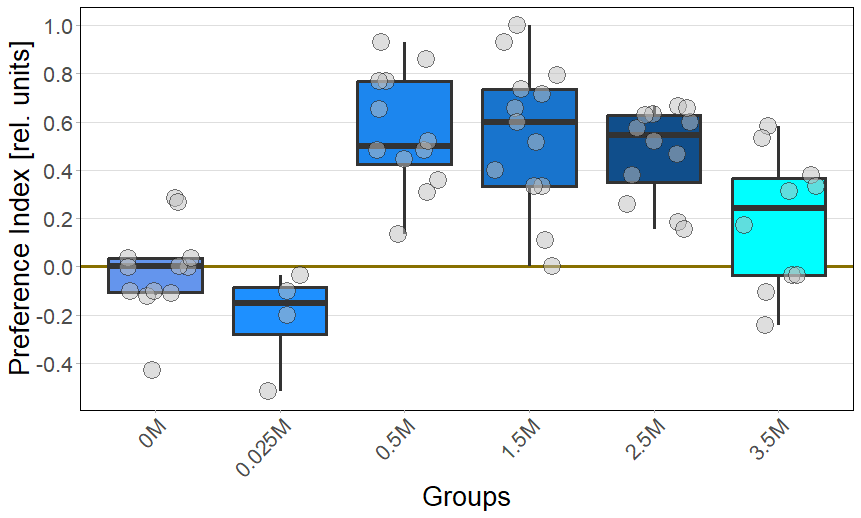
Bluelight
PI: (Pur-Salz)/(Pur+Salz+Neutral)
Stichprobengröße: N(0M) = 12; N(0,025M) = 4; N(0,5) = 12; N(1,5M) = 13; N(2,5M) = 12; N(3,5M) = 10
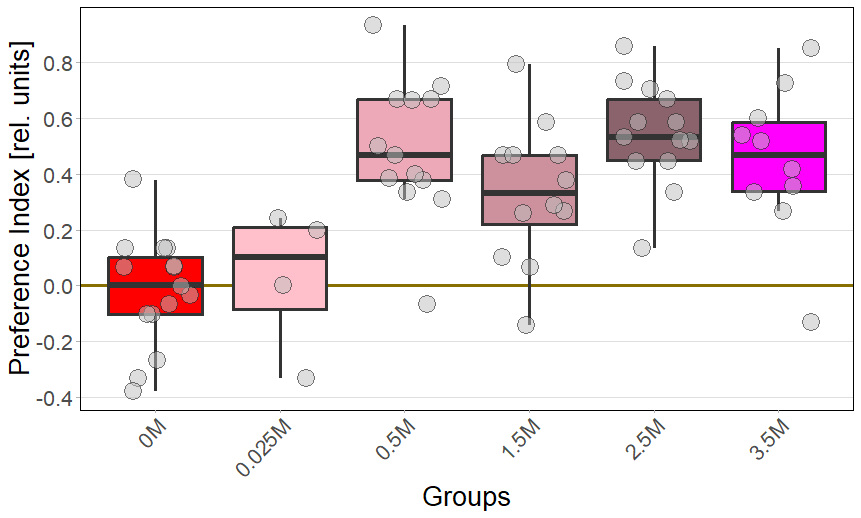
Redlight
PI: (Pur-Salz)/(Pur+Salz+Neutral)
Stichprobengröße: N(0M) = 14; N(0,025M) = 4; N(0,5) = 13; N(1,5M) = 12; N(2,5M) = 13; N(3,5M) = 10
Category: Food preference, Larve, Optogenetics | No Comments
yaw torque measurement_Dtc 20%
on Monday, April 14th, 2025 1:20 | by Julia Schulz
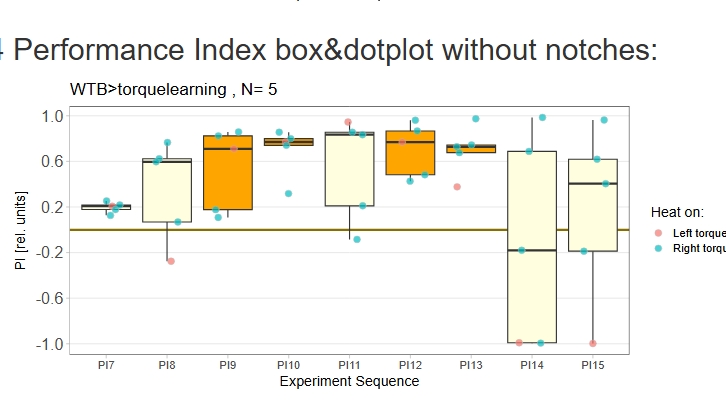
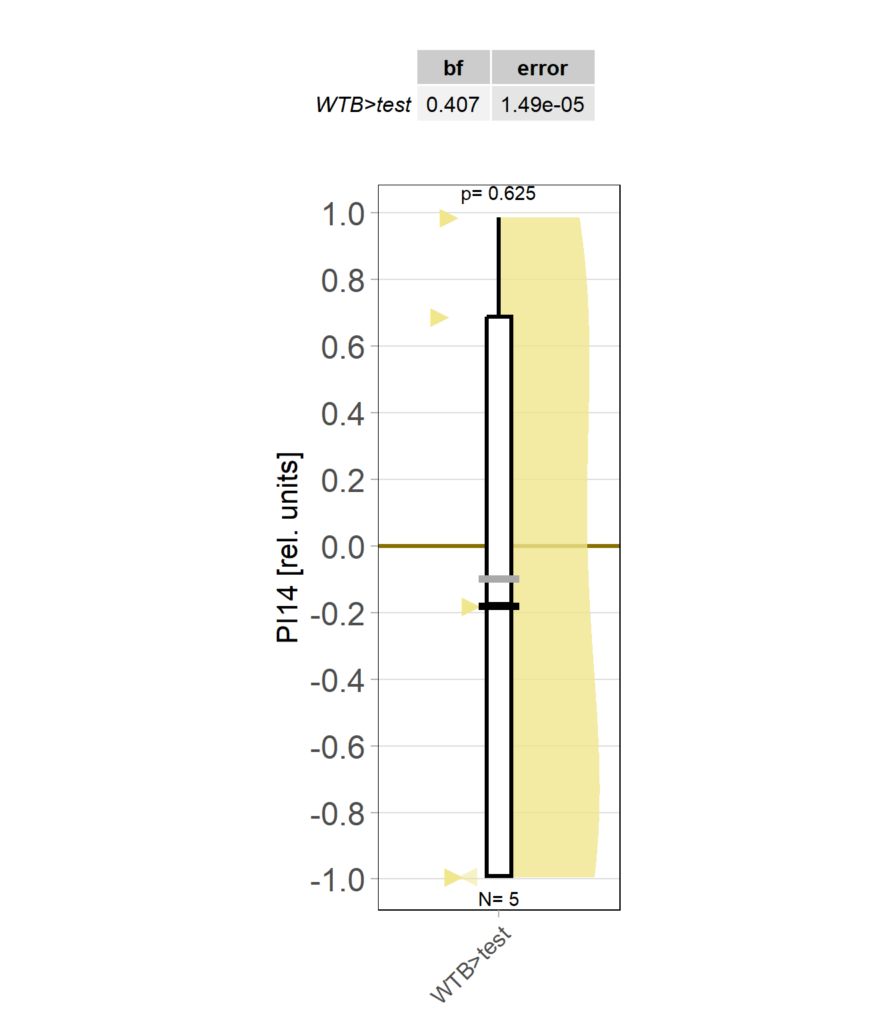
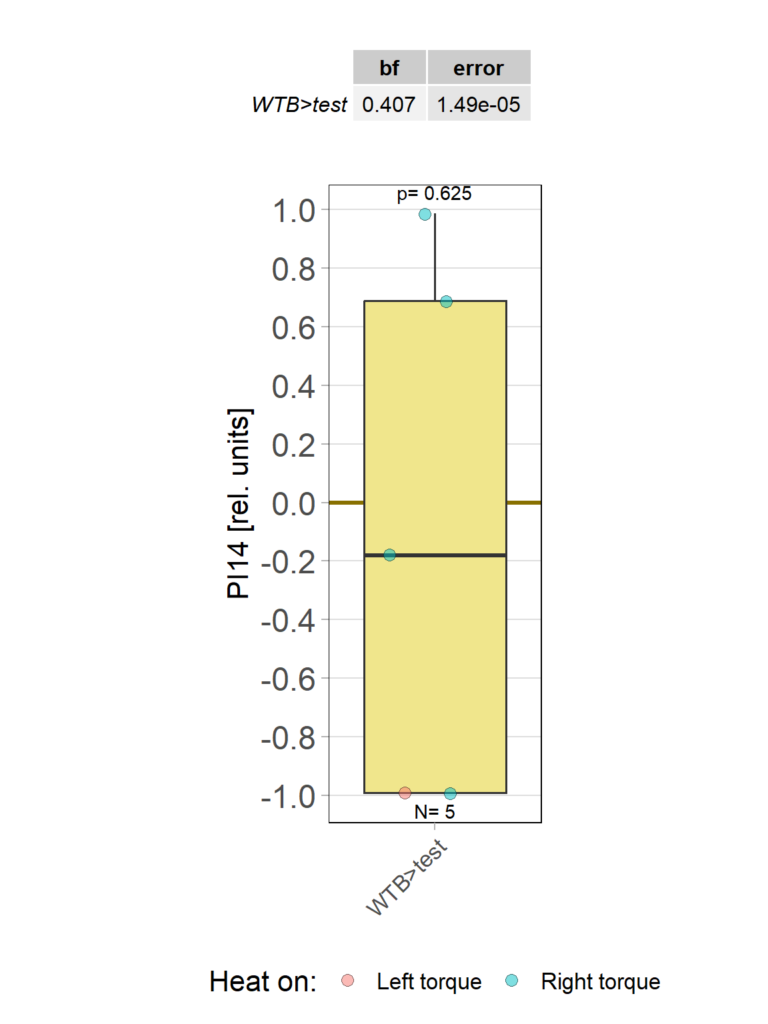
Category: operant self-learning, Optomotor response | No Comments
operant self learning_Dtc 40
on Monday, April 14th, 2025 1:13 | by Julia Schulz
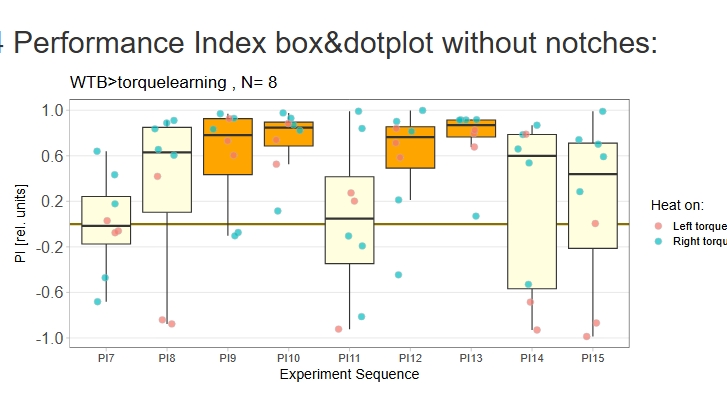
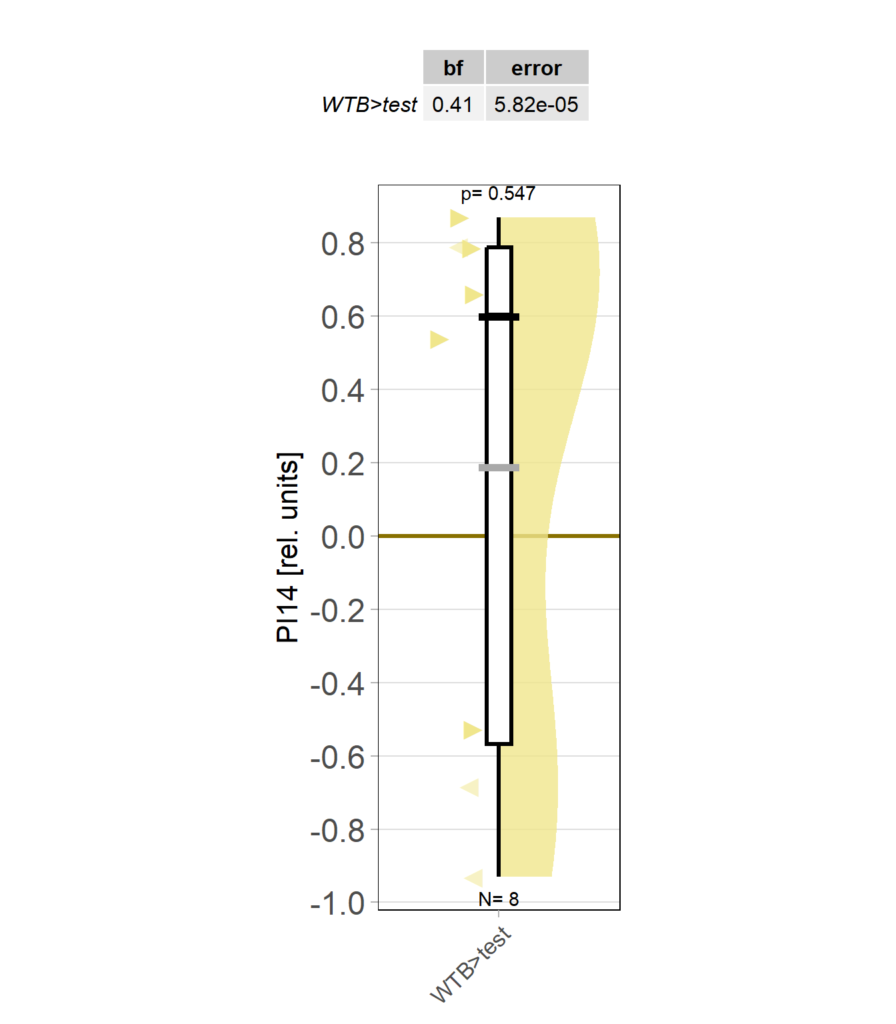
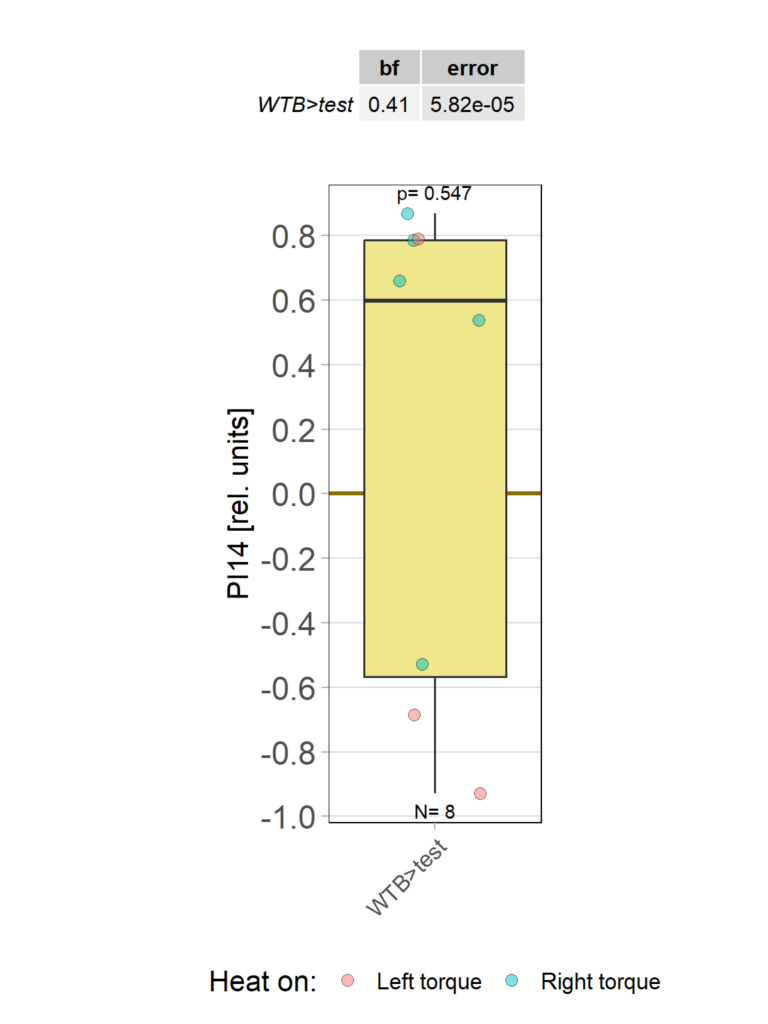
Category: operant self-learning, Optomotor response | No Comments
It’s alive!!
on Thursday, April 10th, 2025 1:31 | by Björn Brembs
Last week, Pavan Kaushik and I finally got the VR-panel setup working. I placed four flies in the machine, one in each VR cube, and then ran each fly three times with a sequence of 8 different pattern wavelengths, also repeated three times, each one rotated both counter-clockwise and clockwise. This is also the sequence which we will use in the course. This is what the data should look like:
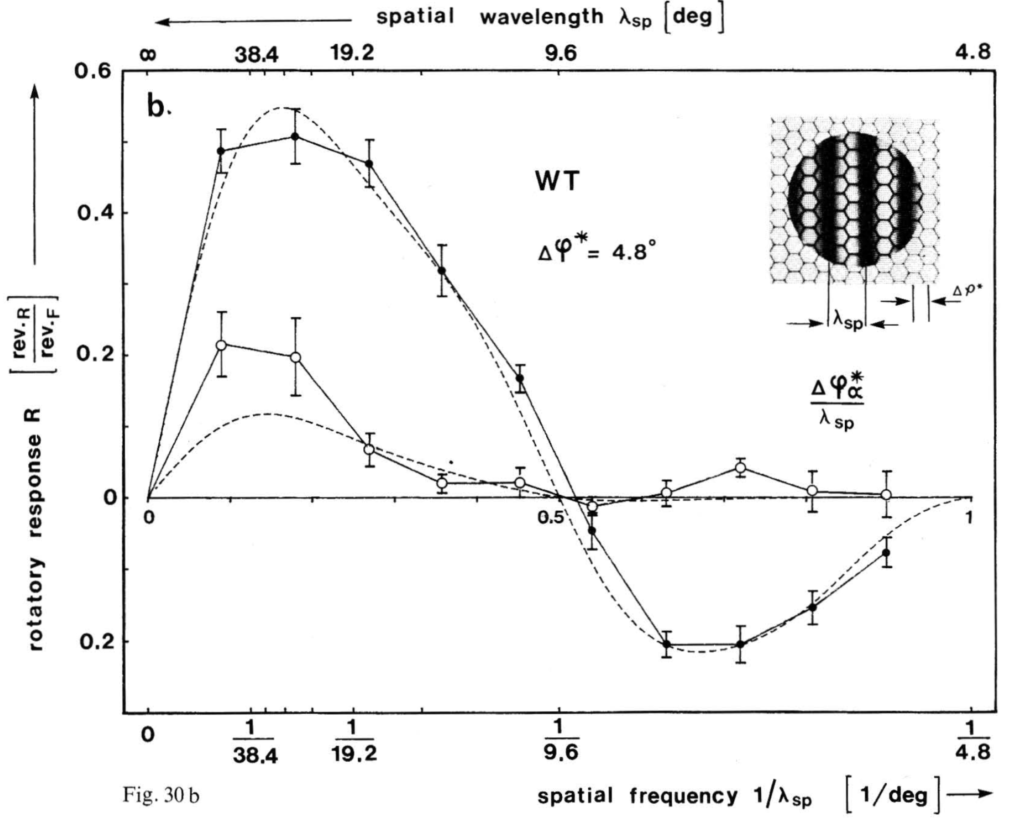
and this is what our data looked like:
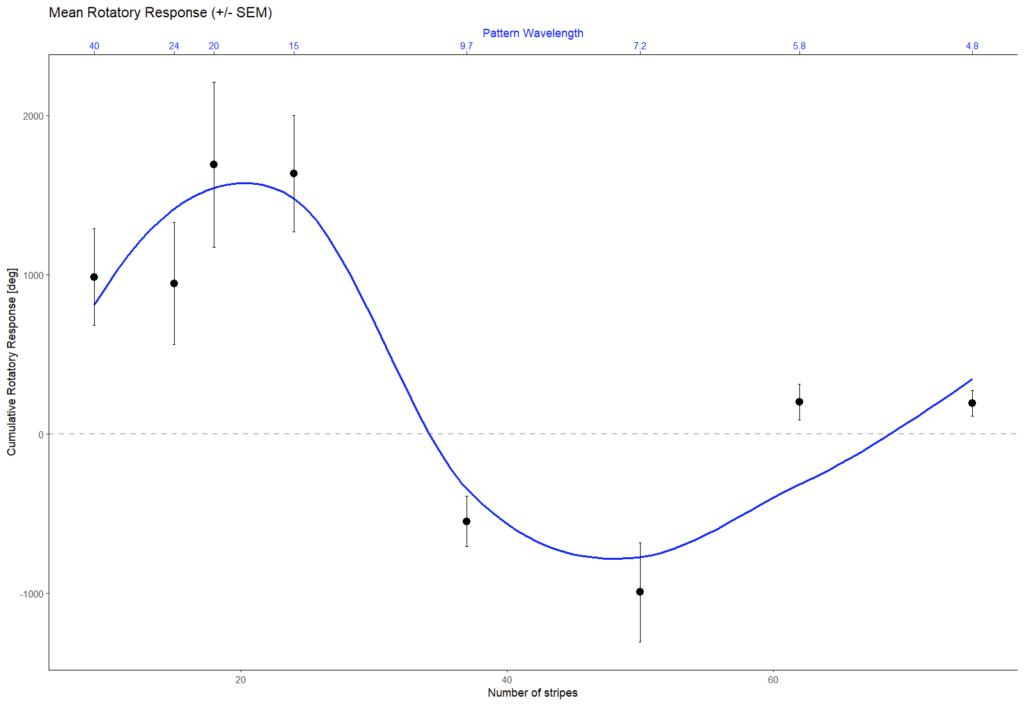
Looks to me like we got it working on the first try. Love that data. Great example to compare the course data against.
The code for evaluating the VR-Panel data is on GitHub.
Category: Optomotor response | No Comments
PPM2 flies avoid the elevator
on Monday, April 7th, 2025 1:14 | by Daniel Döringer
A: Red Light, 1min (N = 25-30)
B: Yellow Light, 1min (N = 26-30)
C: Red Light, 10 min (N = 18-30)
D: Yellow Light, 10min (N = 16-30)
Category: Uncategorized | No Comments
Salt Avoidance CantonS
on Monday, April 7th, 2025 12:48 | by Eva Schächtl

Stichprobengröße pro Gruppe: N = 10
PI: (Pur-Salz)/(Pur+Salz+Neutral)

Stichprobengröße pro Gruppe: N = 10 und bei 0M N = 15
PI:(Pur-Salz)/(Pur+Salz+Neutral)
Category: Food preference, Lab, Larve, Optogenetics | No Comments
yaw torque learning_Dtc_40%
on Monday, March 31st, 2025 10:30 | by Julia Schulz
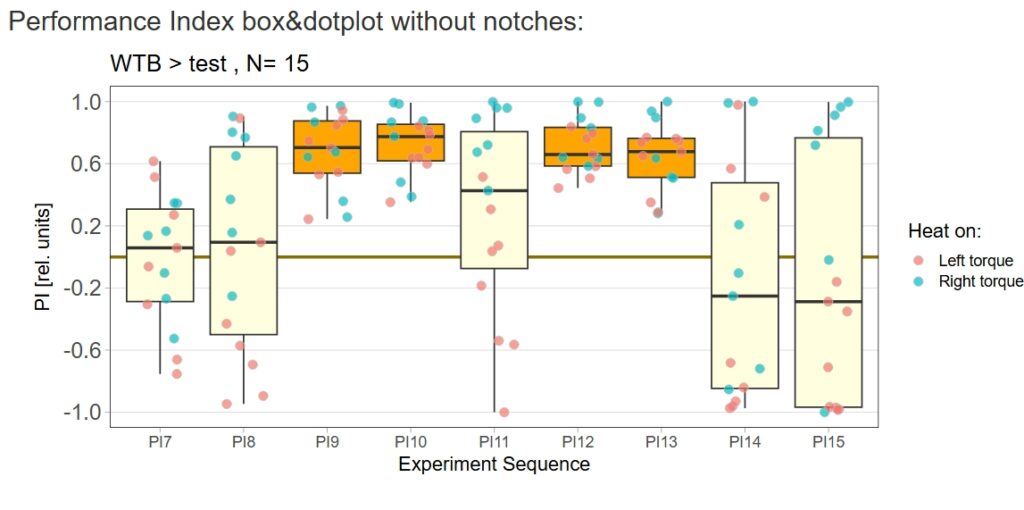
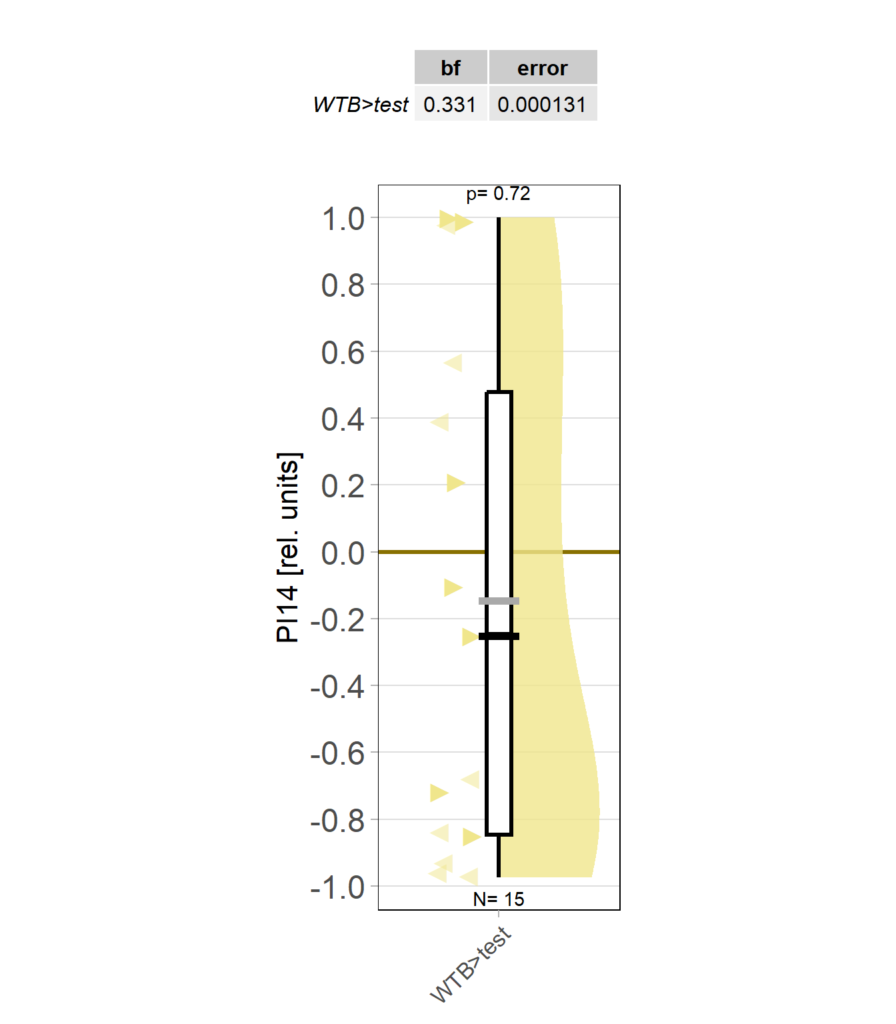
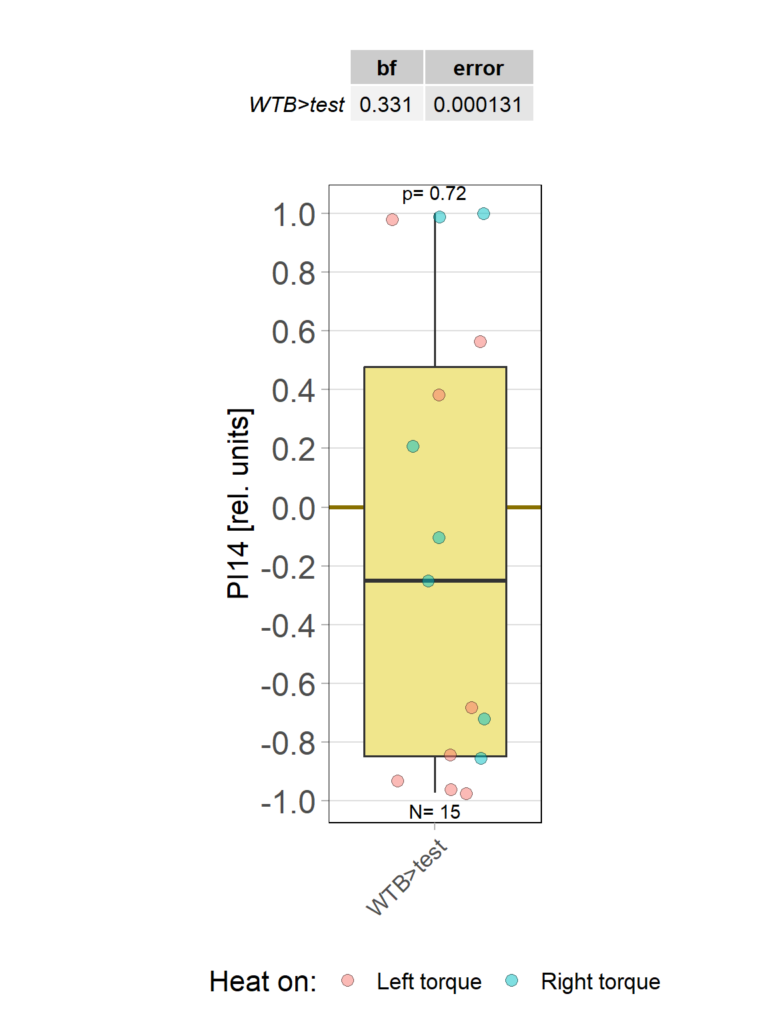
Category: Operant learning, operant self-learning | No Comments
Not an unambiguous result
on Friday, March 28th, 2025 3:53 | by Björn Brembs
This is the final graph of the experiments knocking down PKG (foraging) in operant self-learning:

The data have been automatically uploaded via the new script and can be found on the University’s publication server.
This was supposed to be a reproduction of Tina’s experiment:

While in my dataset, the knock-down flies are still performing the worst of all three groups, it is not quite as clear-cut as in Tina’s results. Then again, her knock-down flies maybe were slightly too low. Now pooled with mine that were slightly to high, maybe this is what we should expect: a clear zero. We should pool the data and see how it looks.
Category: operant self-learning, PKG | No Comments
Generation of UAS-aPKC-TurboID lines
on Monday, March 24th, 2025 1:44 | by Julia Schulz
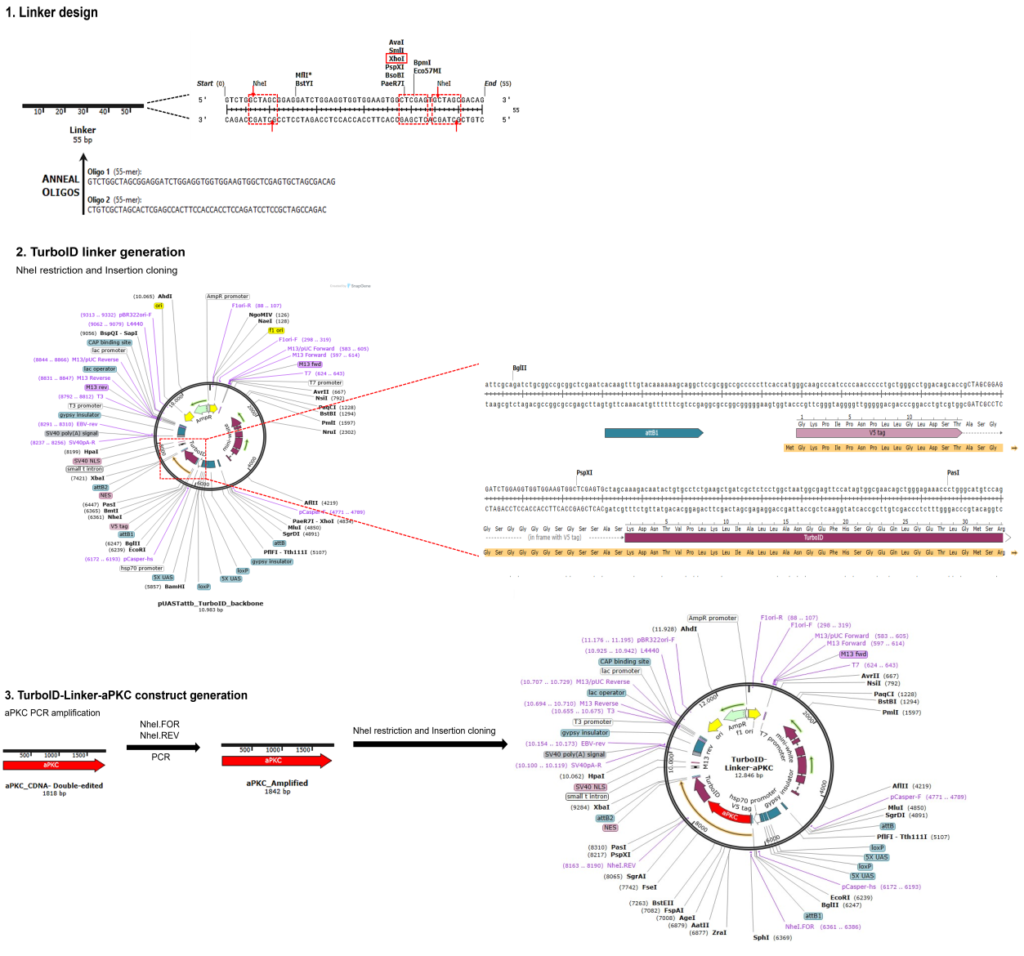
Category: operant self-learning, PKC | No Comments
Salt avoidance under blue light canton S
on Monday, March 24th, 2025 1:08 | by Eva Schächtl
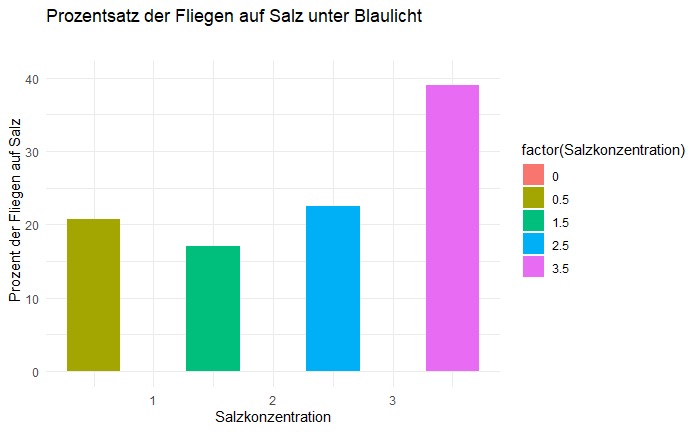
Category: Food preference, Lab, Larve | No Comments
