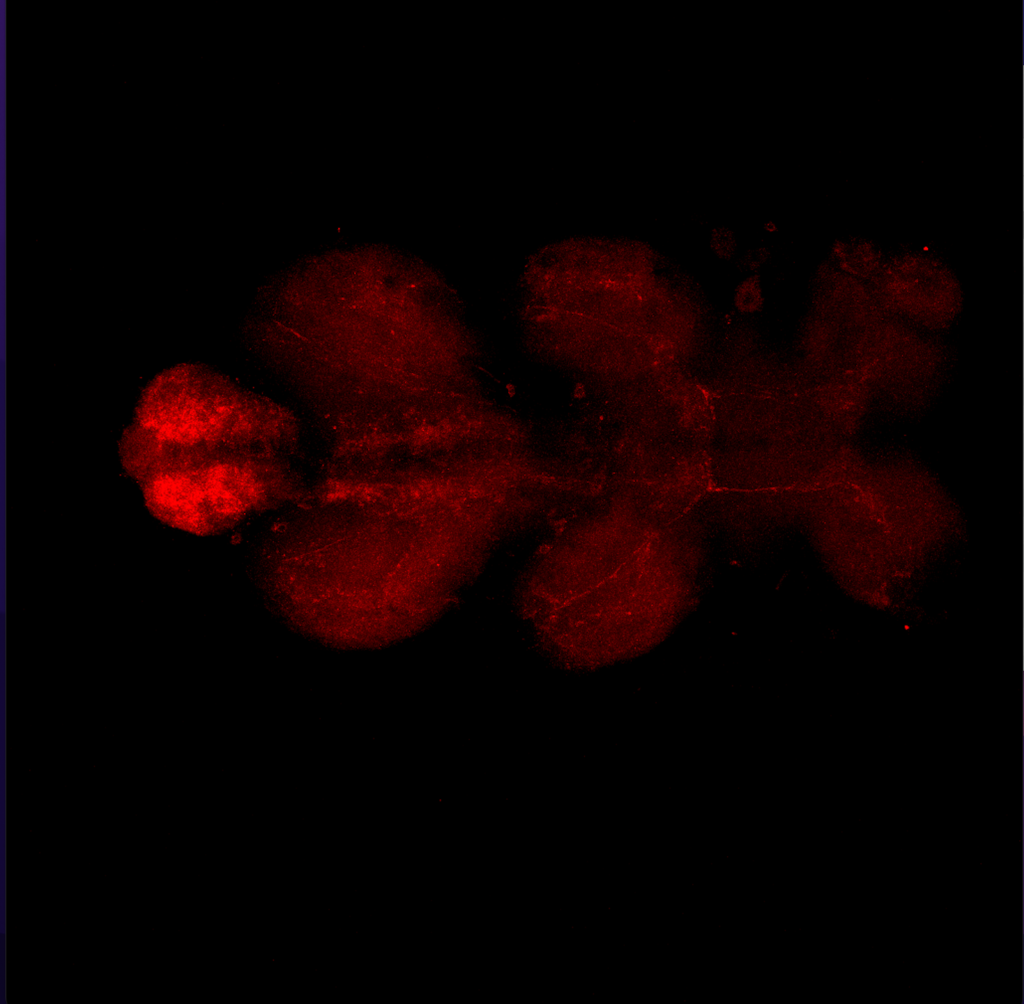
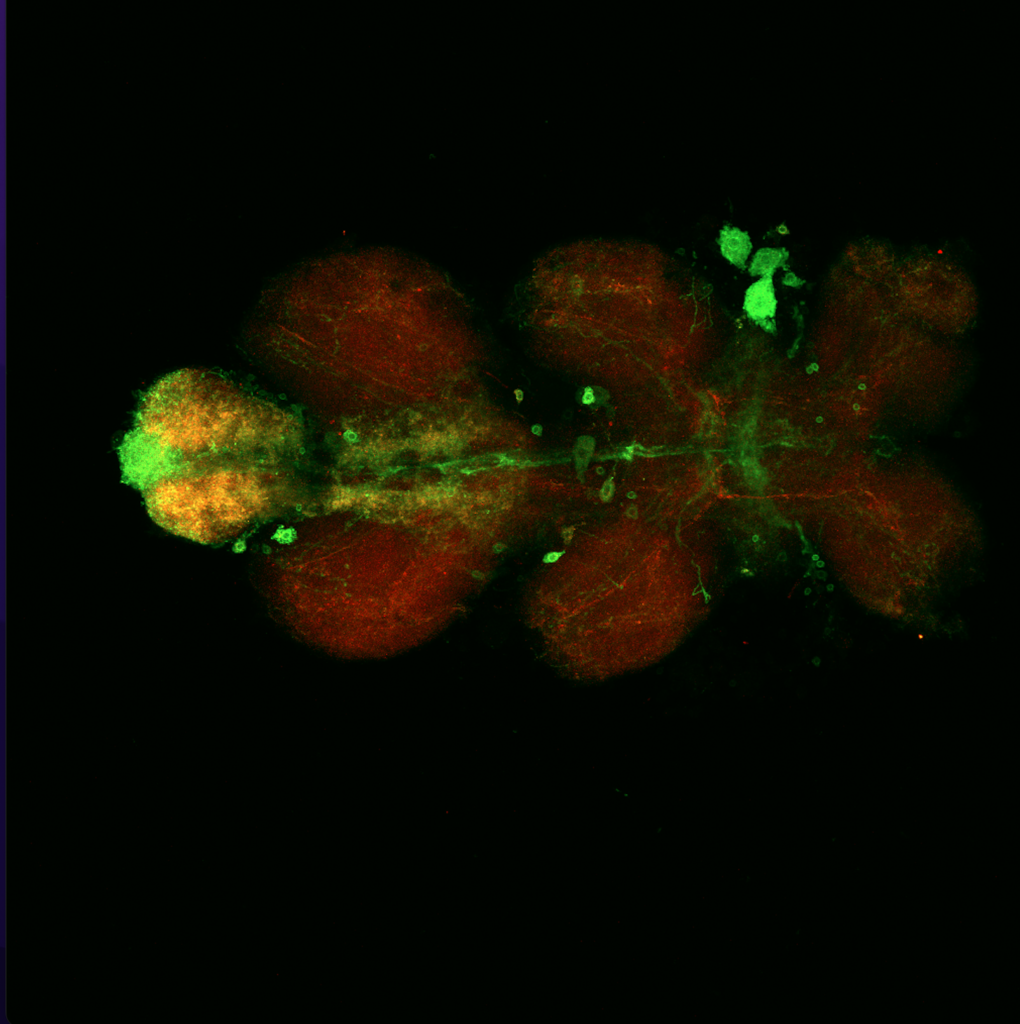
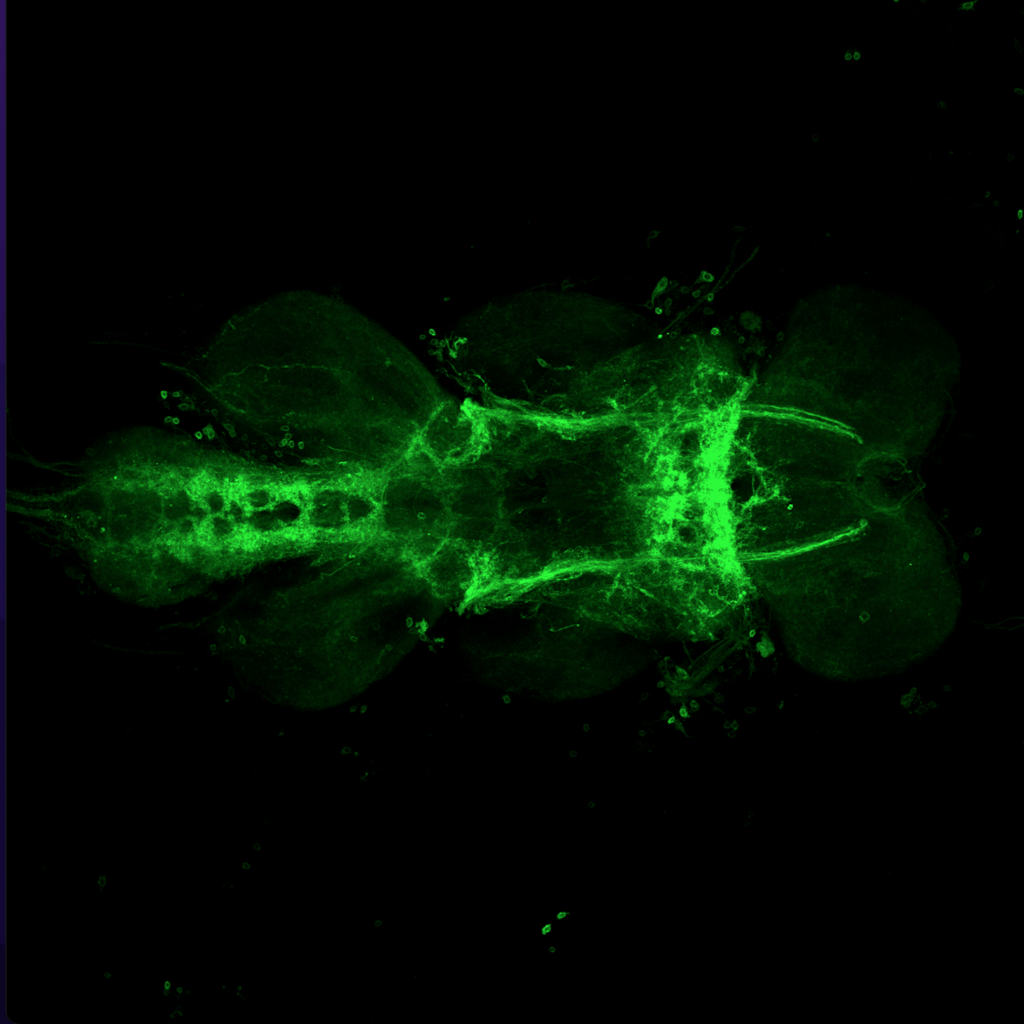
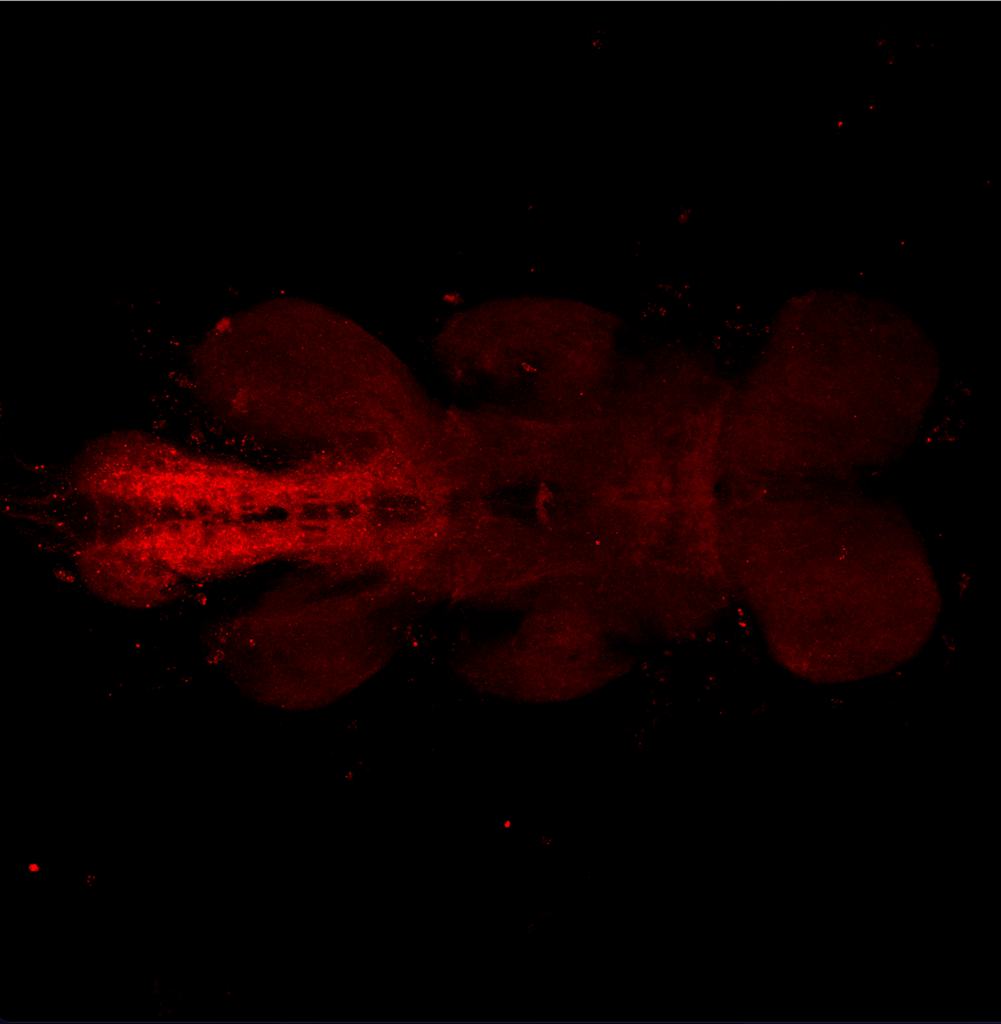
Results for MB+DANs and MB054B
on Monday, September 25th, 2023 11:07 | by Maja Achatz
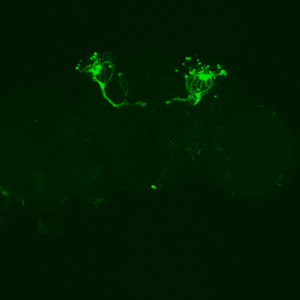
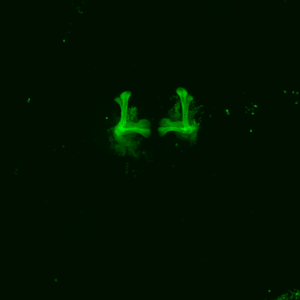
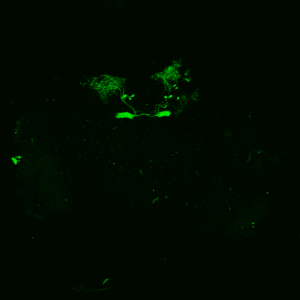
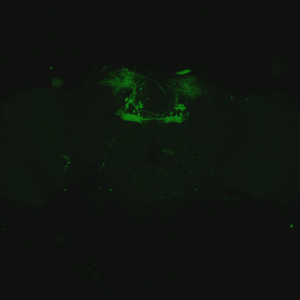
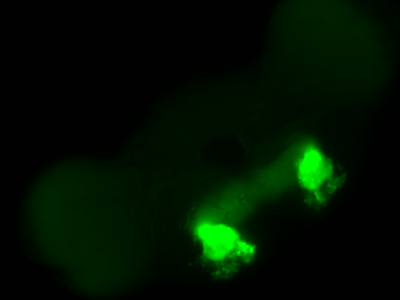
The first experiment tested the activation of KCs with simultaneous ablation of pPAM which showed the following results:
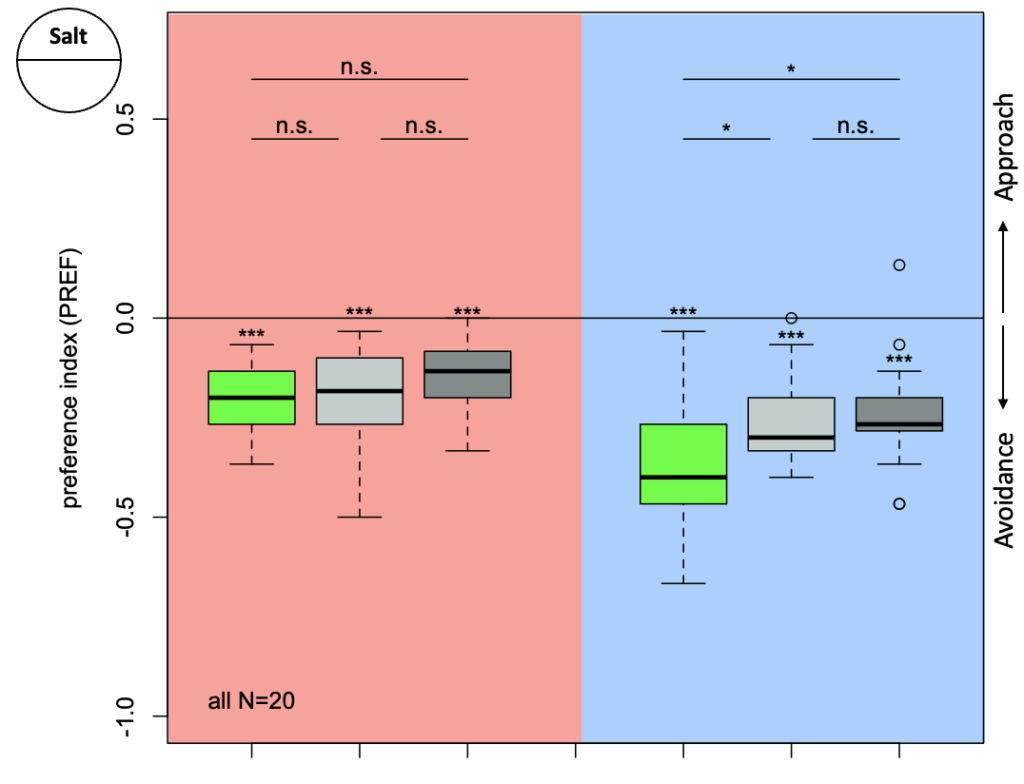
Green: UAS-ChR2-XXL; LexAop-rpr x 58E02-LexA; H24-Gal4
Lightgrey: UAS-ChR2-XXL; LexAop-rpr x H24-Gal4
Darkgrey: UAS-ChR2-XXL x 58E02-LexA; H24-Gal4
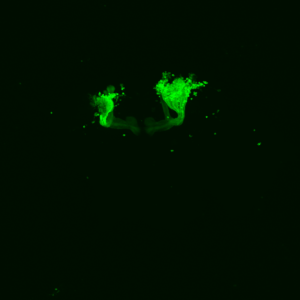
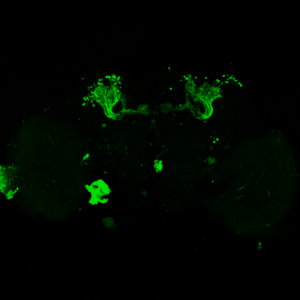
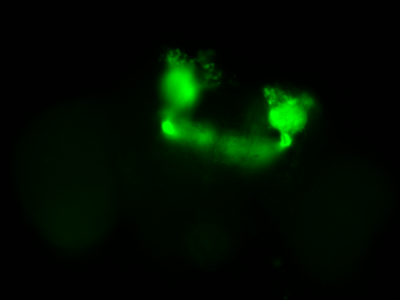
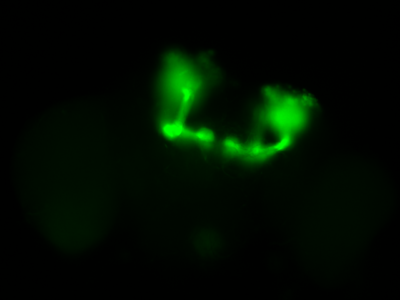
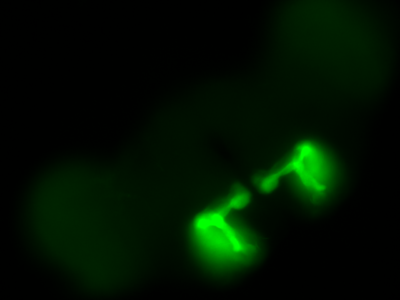
The next experiment was conducted with the split-Gal4 line MB054B (DAN-f1/DAN-g1) and showed the following:
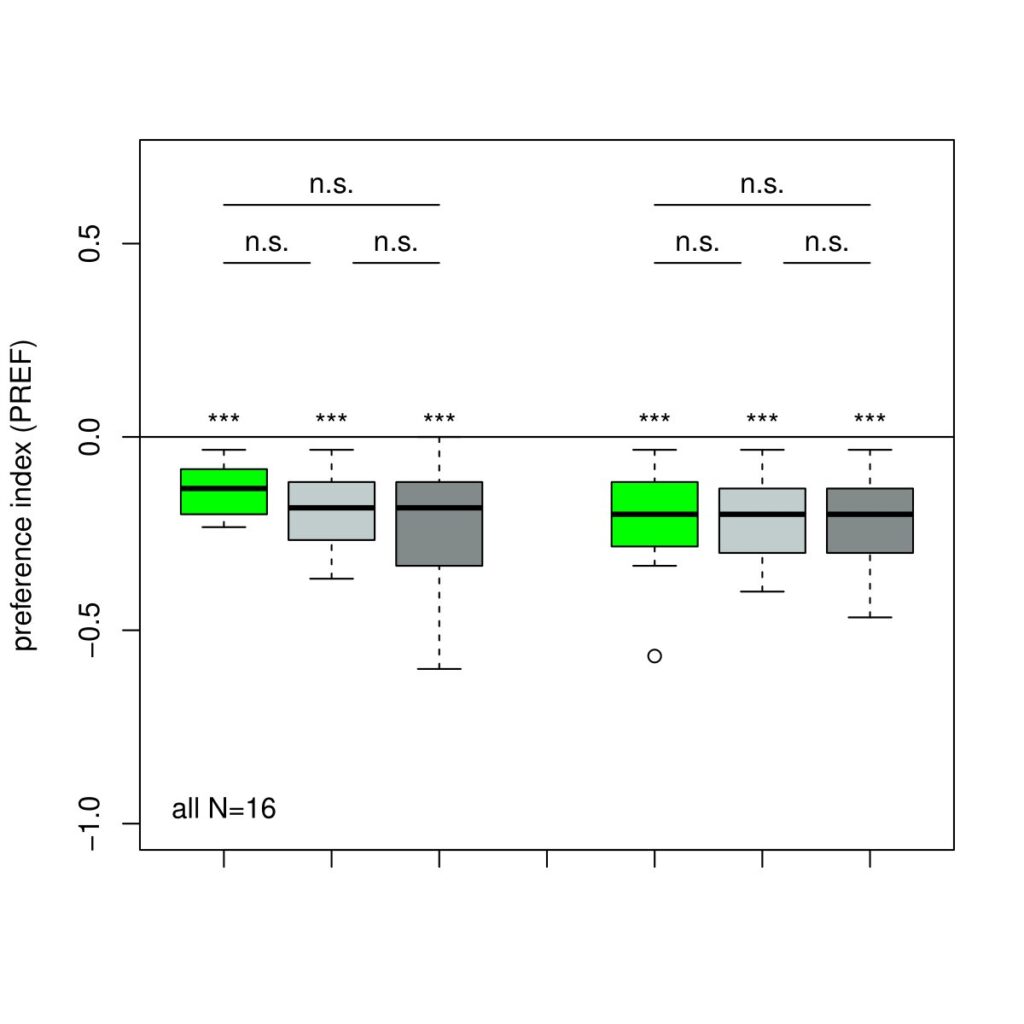
Category: Kenyon cells, Larve, Mushroom Body, Optogenetics | No Comments
Verification TH-C-AD;TH-D-DBD
on Monday, September 11th, 2023 9:52 | by Luisa Guyton
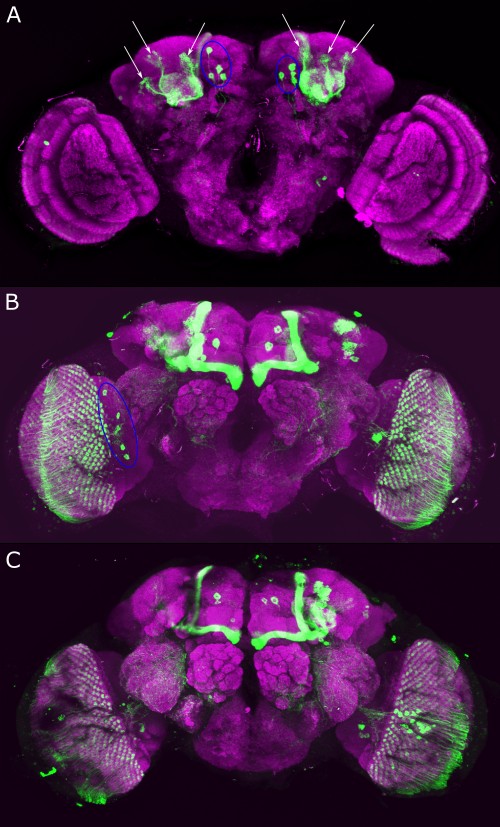
Category: Anatomy, DAN, genetics, Kenyon cells, Optogenetics | No Comments
Line verification SS56699
on Tuesday, August 29th, 2023 7:50 | by Luisa Guyton
After dissecting brains from the GAL4 driver line SS56699 with GFP staining and finding no fluorescence, I dissected them again and the immunohistochemical staining showed fluorescence. To check that the correct neurons were stained, I compared the images with the image in the paper by Hulse et al (2021; https://doi.org/10.7554/eLife.66039) . The three PPL1 dopaminergic dorsal fan-shaped body tangential neurons per hemisphere are stained, but there are some additional unidentified neurons (presumably PPM1 neruons) visible.
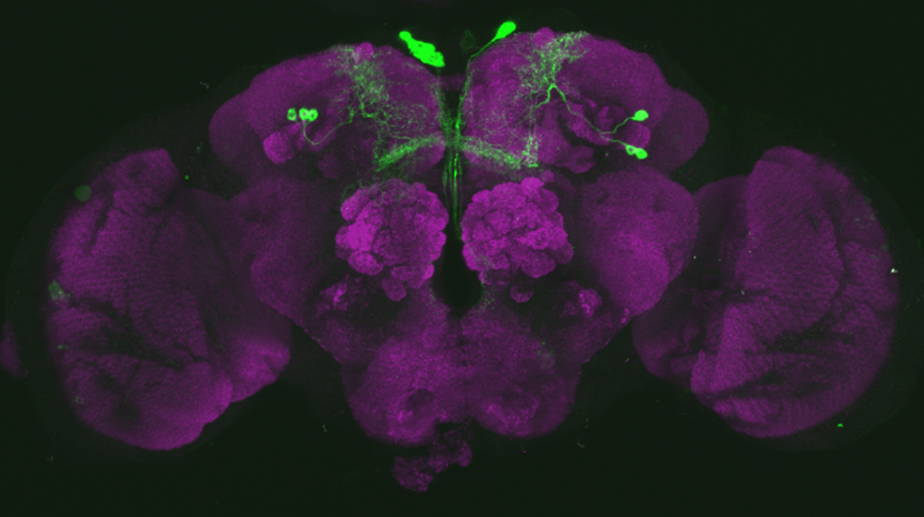
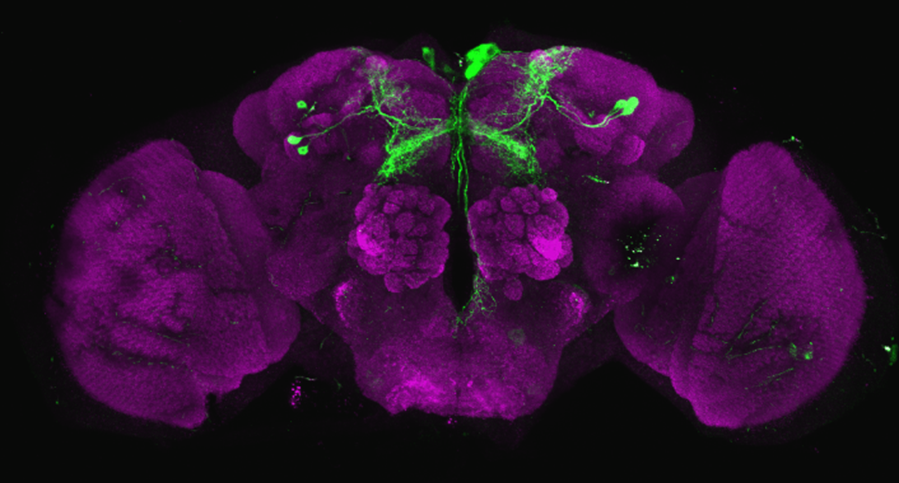
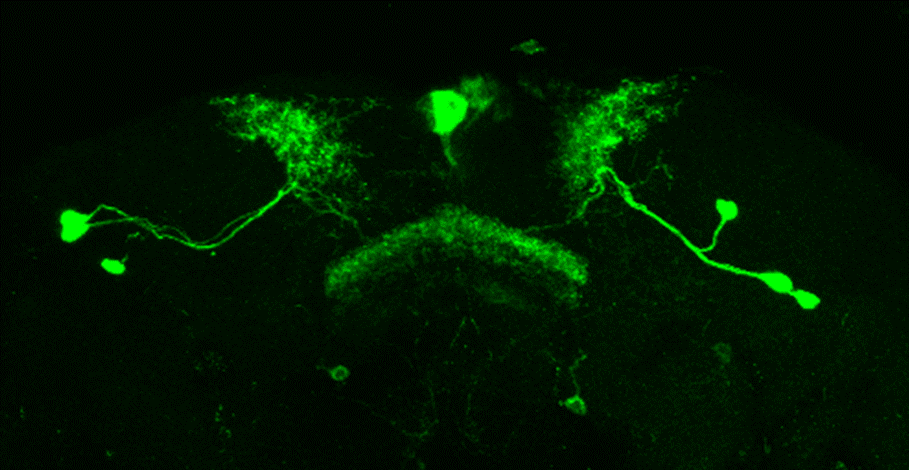
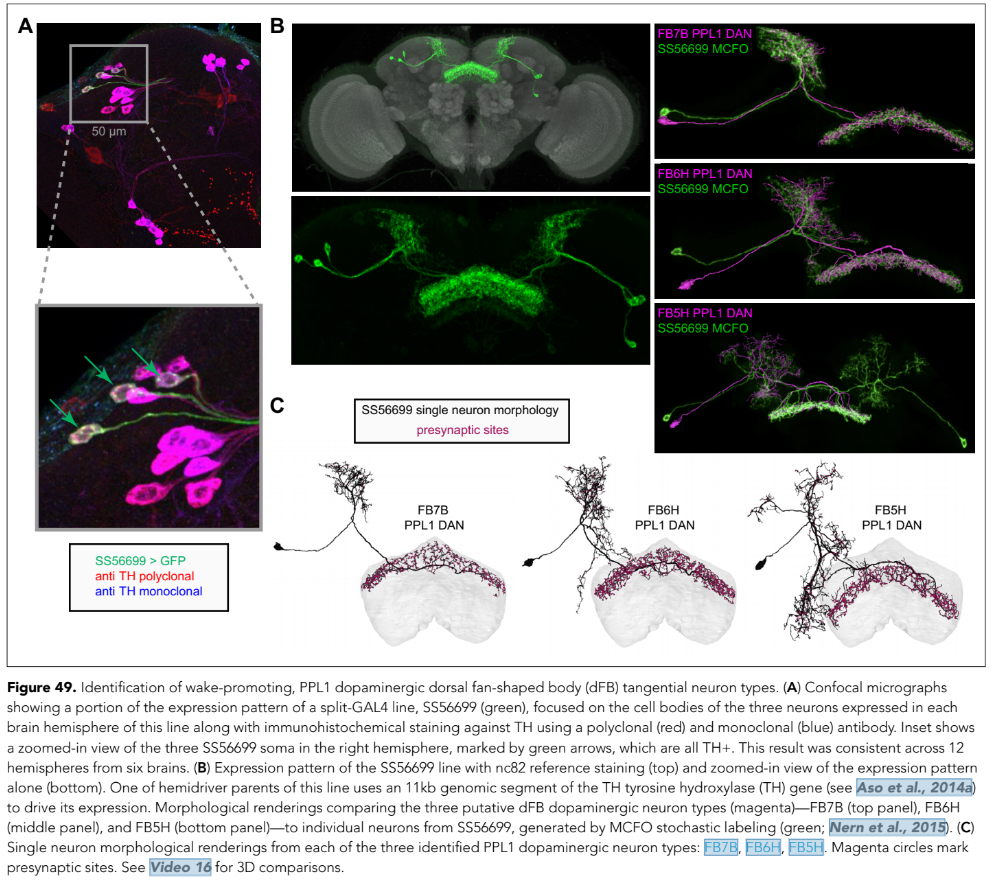
Hulse, B. K., Haberkern, H., Franconville, R., Turner-Evans, D., Takemura, S.-y., Wolff, T., Noorman, M., Dreher, M., Dan, C., Parekh, R., Hermundstad, A. M., Rubin, G. M., & Jayaraman, V. (2021). A connectome of the Drosophila central complex reveals network motifs suitable for flexible navigation and context-dependent action selection. eLife, 10, e66039. https://doi.org/10.7554/eLife.66039
Category: Anatomy, genetics | No Comments
Lines Verification
on Monday, September 12th, 2022 9:55 | by Silvia Marcato
Category: Anatomy, DAN, Kenyon cells, Mushroom Body, PAM | No Comments
MB Lines verification
on Monday, July 18th, 2022 11:11 | by Radostina Lyutova
Category: Anatomy, Mushroom Body | No Comments
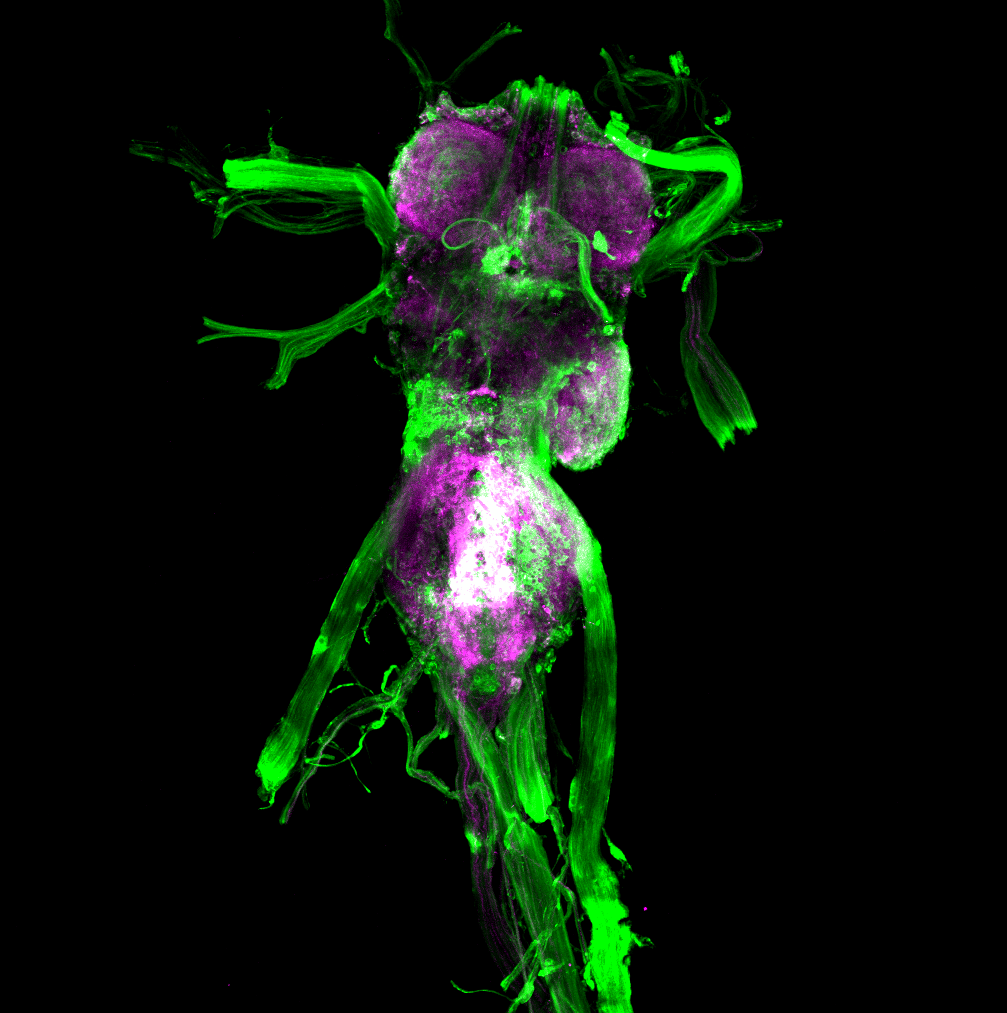
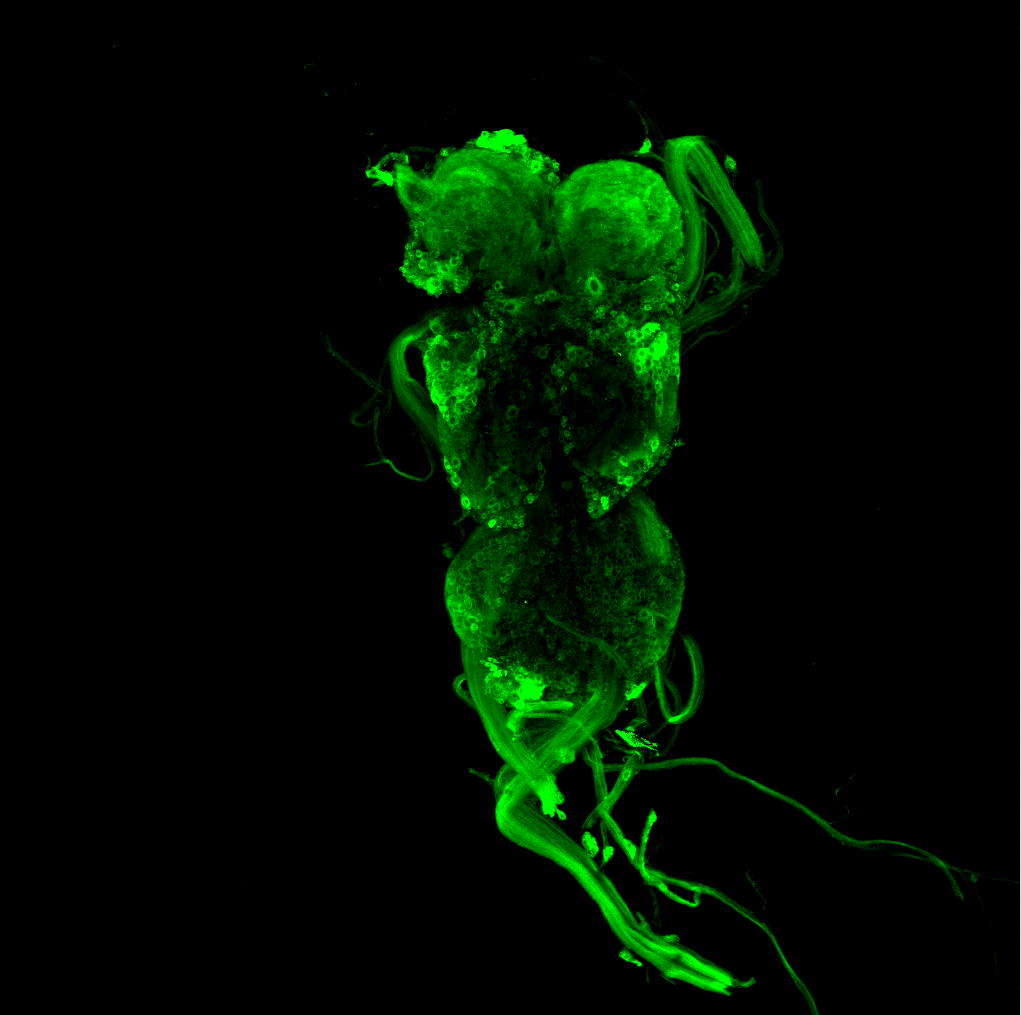
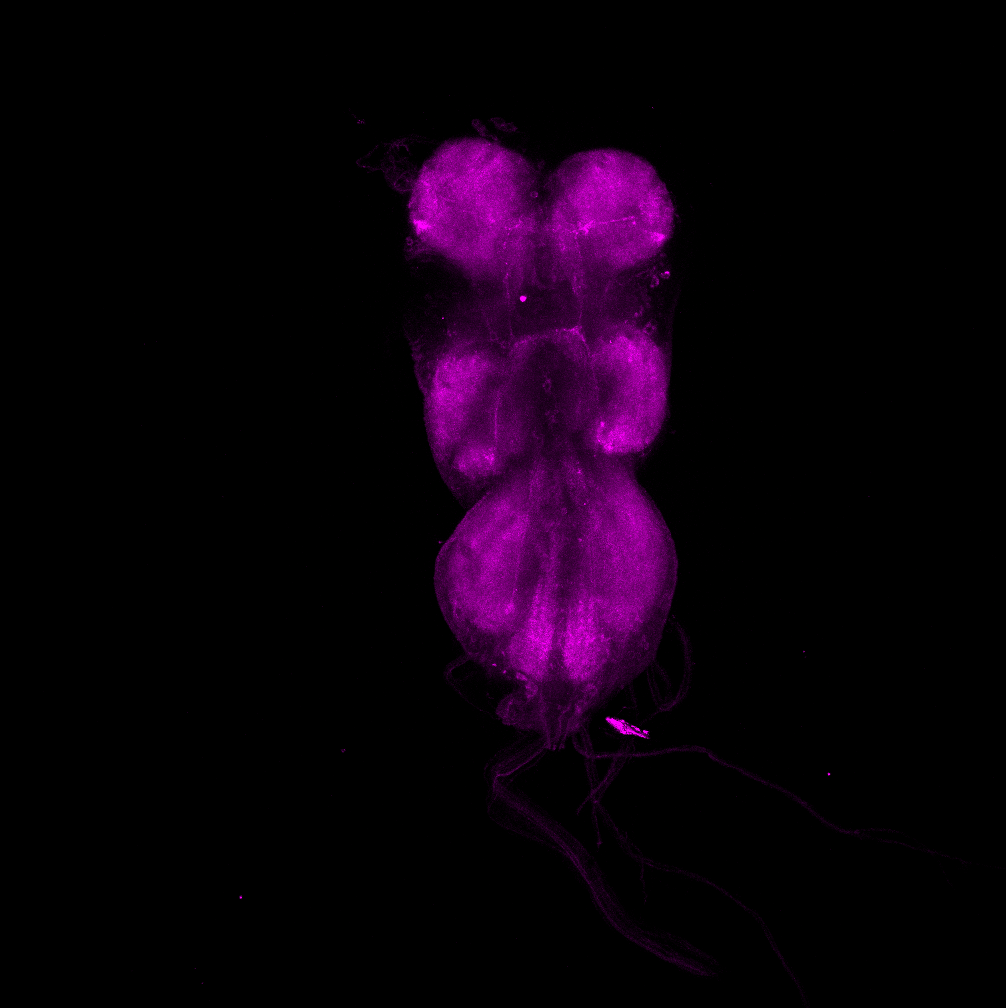
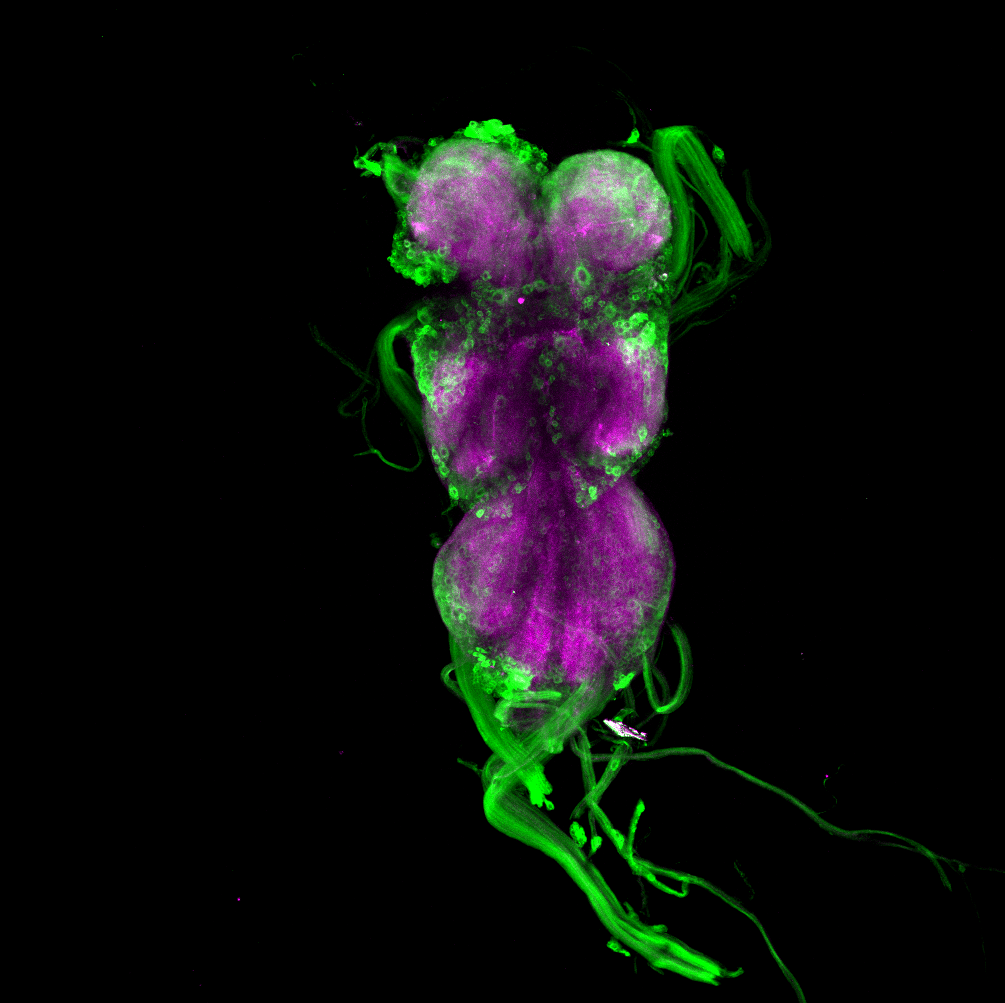
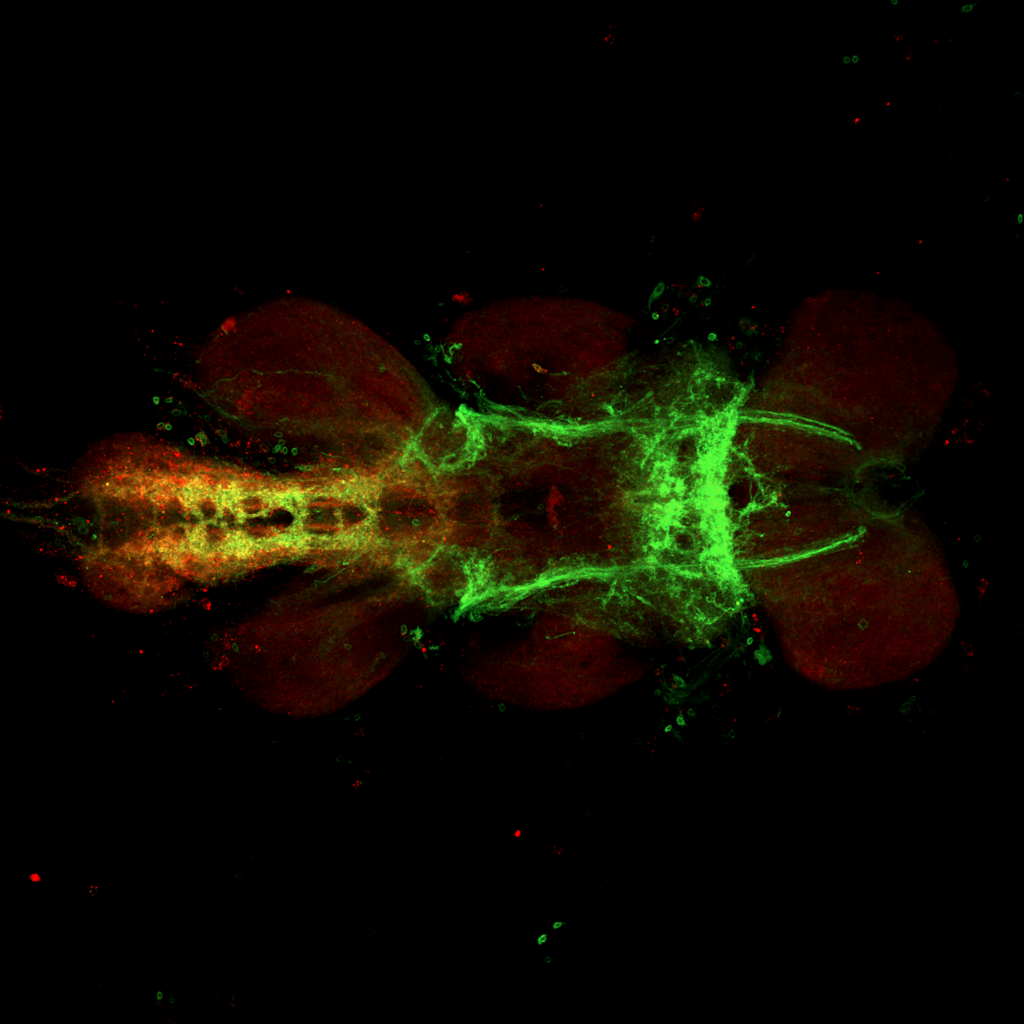
Overlap of aPKC and FoxP in the VNC projecting to the wings
on Monday, June 27th, 2022 12:54 | by Amelie Hauser
Green: aPKC (GFP)
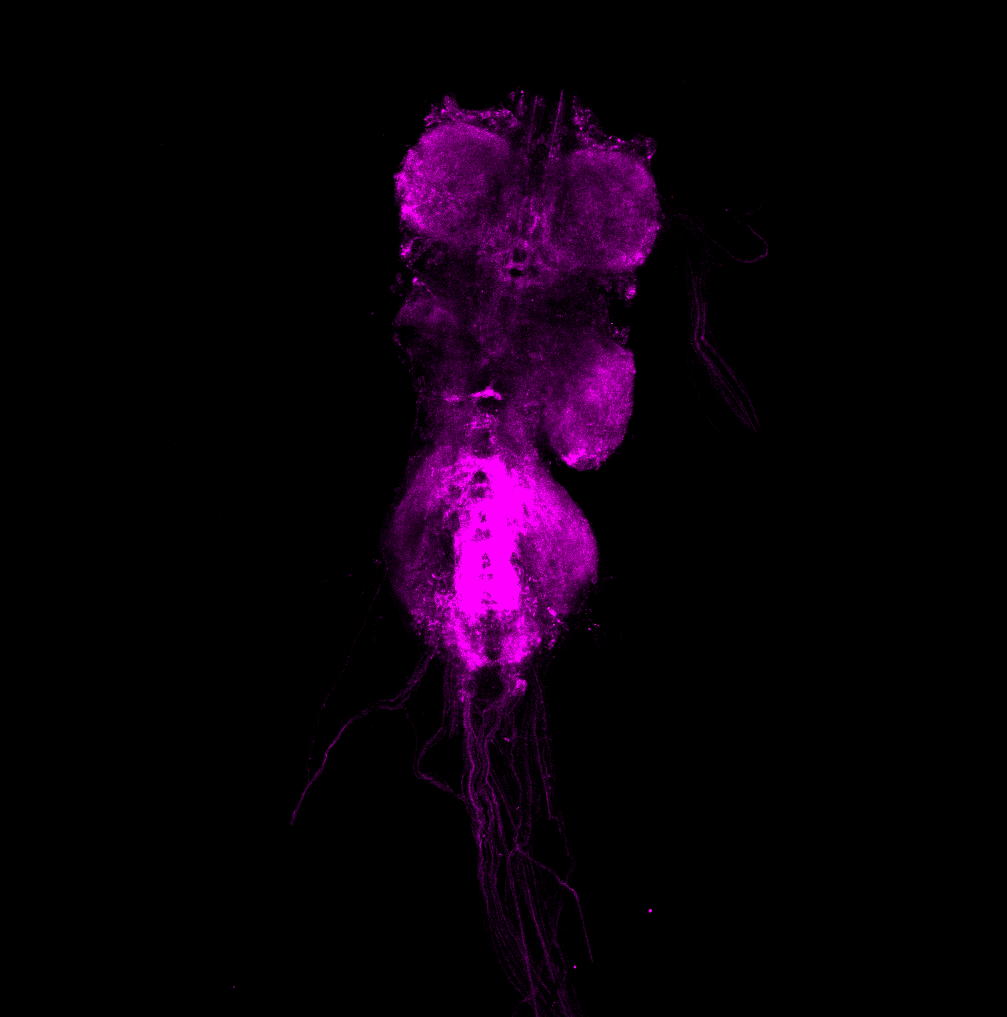
Magenta: FoxP (RFP)
White: Overlap marked with the blue circle projecting to the wings
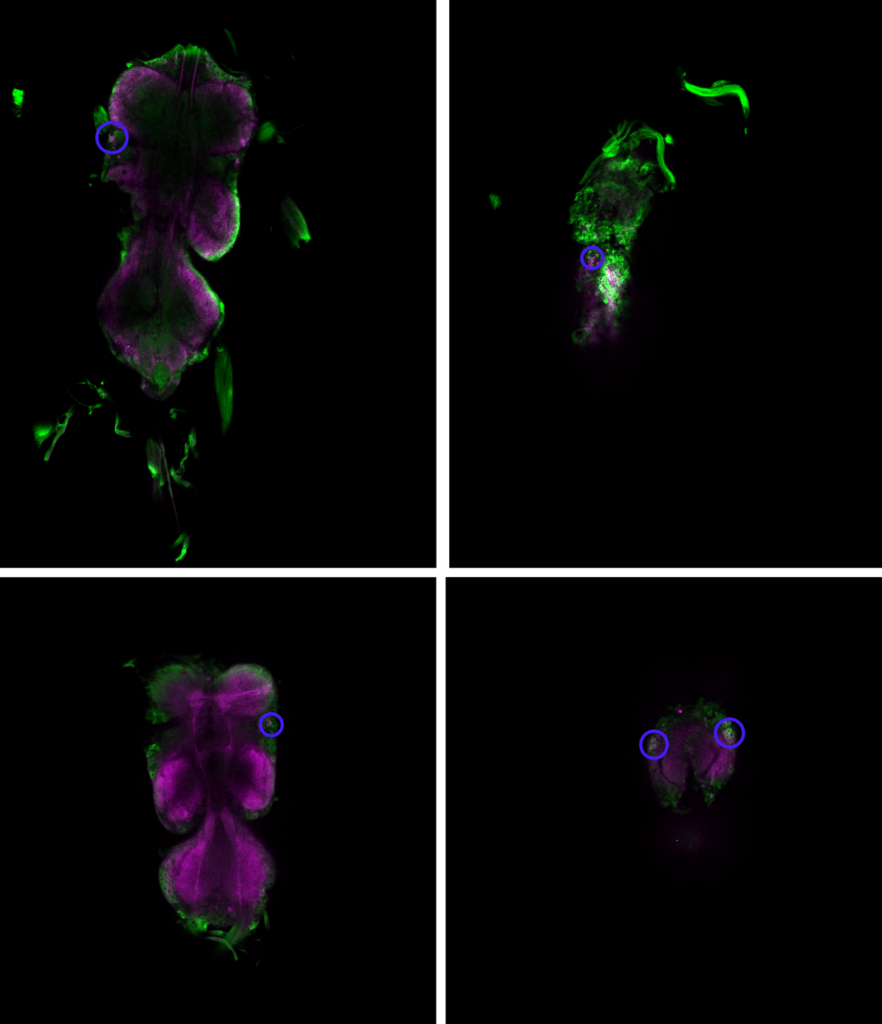
Category: Anatomy | No Comments
aPKC/FoxP expression in the VNC with antibody staining; dorsal view
on Sunday, June 26th, 2022 10:52 | by Amelie Hauser
Category: Anatomy | No Comments
aPKC/FoxP expression in the VNC using antibody staining; ventral view
on Sunday, June 26th, 2022 10:50 | by Amelie Hauser
Category: Anatomy | No Comments
Anatomy comparison OK6/FoxP
on Monday, May 23rd, 2022 1:45 | by Amelie Hauser
Category: Anatomy | No Comments
Anatomy comparison D42/FoxP
on Monday, May 23rd, 2022 1:41 | by Amelie Hauser
Category: Anatomy | No Comments