Habit Formation Wildtype Test 02.03-06.03
I wanted to try Habit formation with wildtype and tested 4 wheels of flies (48) over the last week. Unfortunately, I did a mistake when setting the timeslots file for right torque so it had an additional testing period at P13. So analyzed the left and right individually:
For Punishment on Left Torque:
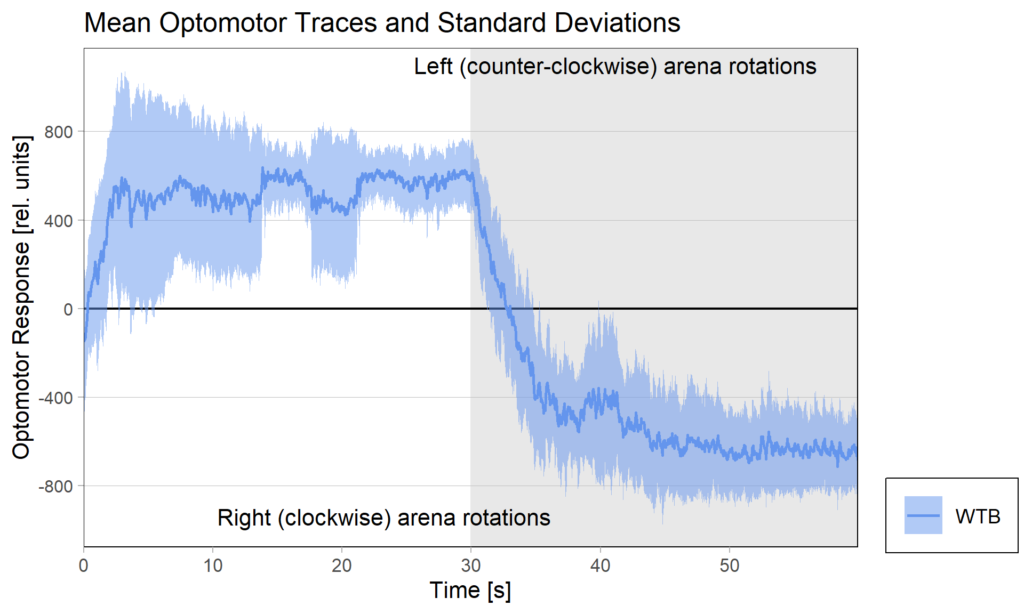
Optomotor traces (right/left) at start of experiment:
Performance Index box&dotplot without notches:
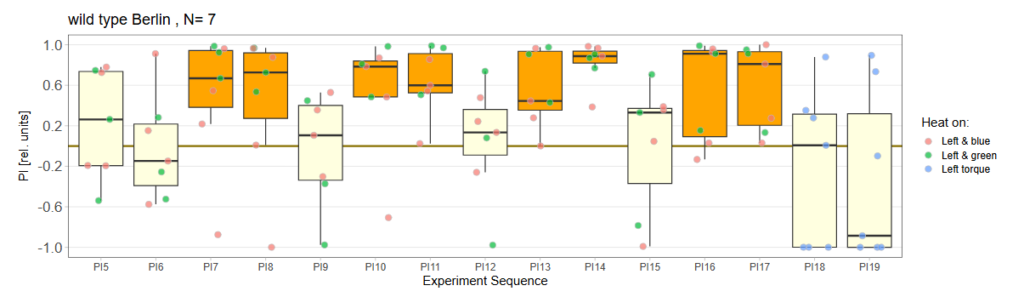
For Punishment on Right Torque:
Optomotor traces (right/left) at start of experiment:
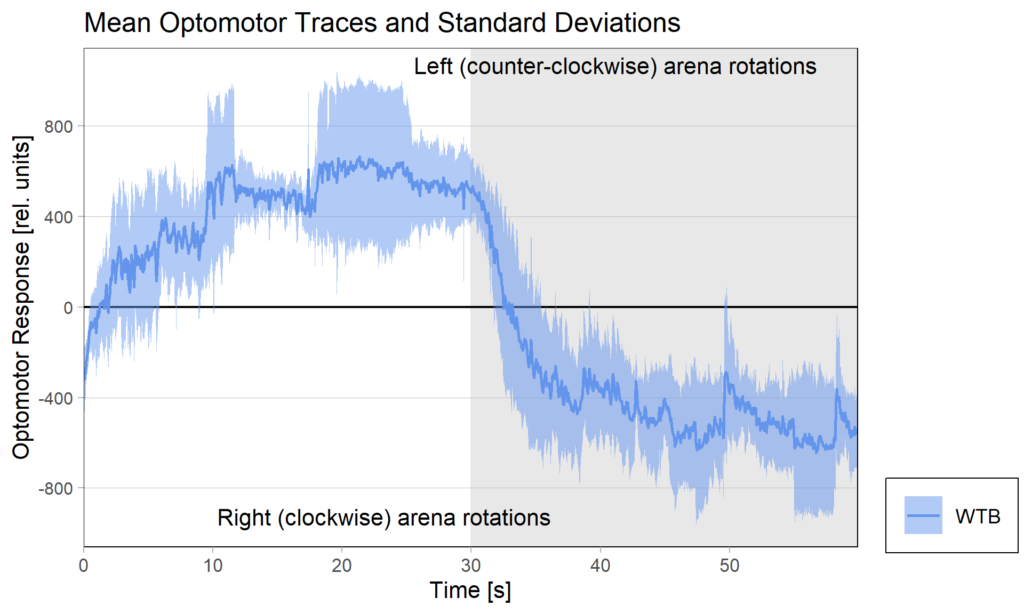
Performance Index box&dotplot without notches:
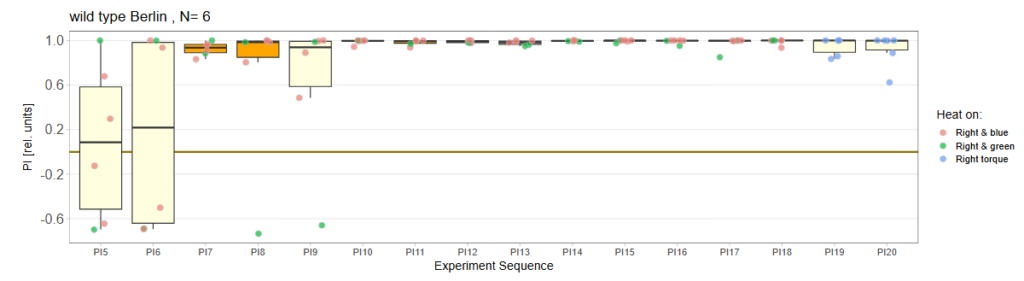
WTB Test for Torquemeter February 19-20
10 flies completed out of 36 flies.
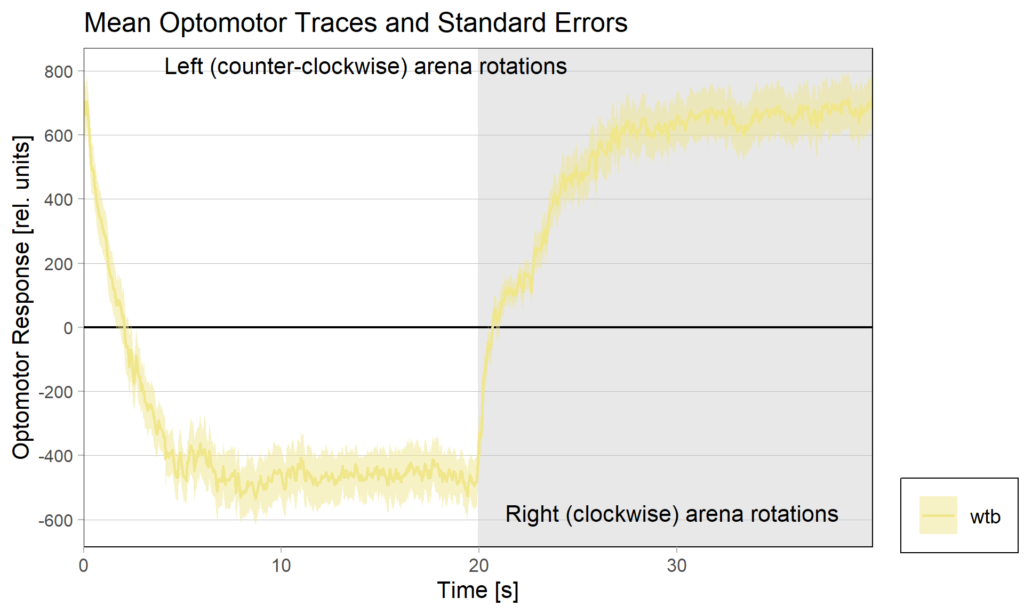
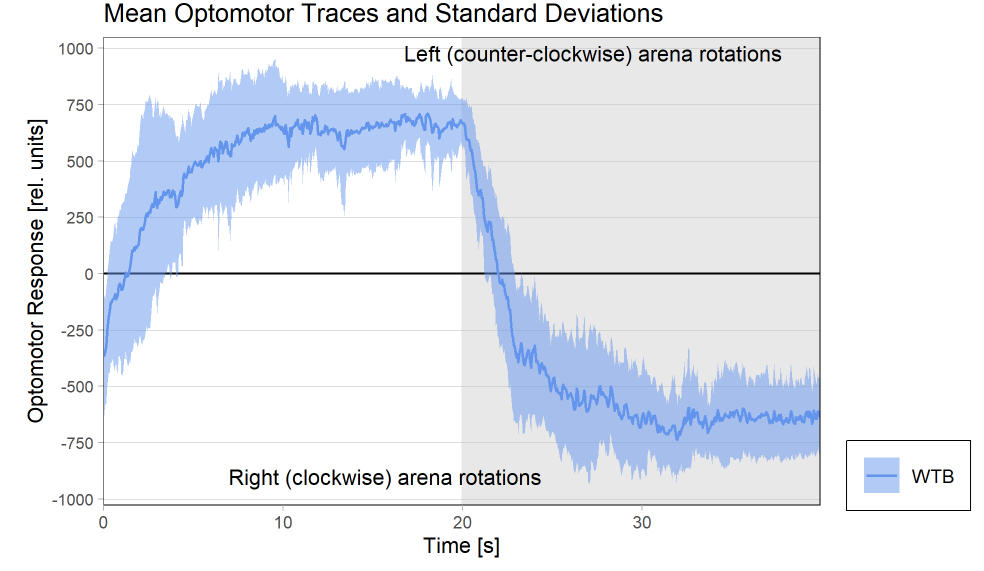
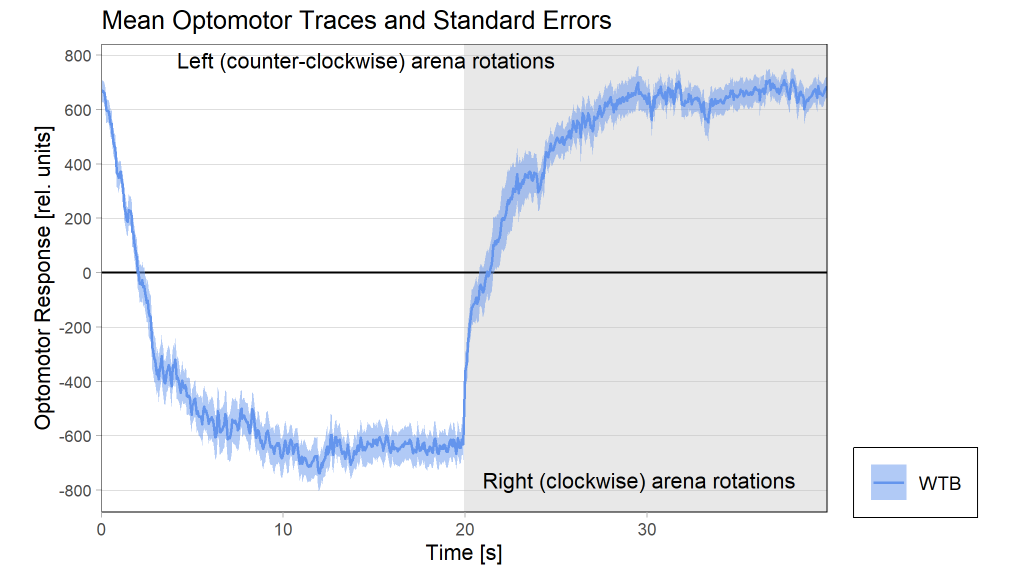
Optomotor traces (right/left) at start of experiment
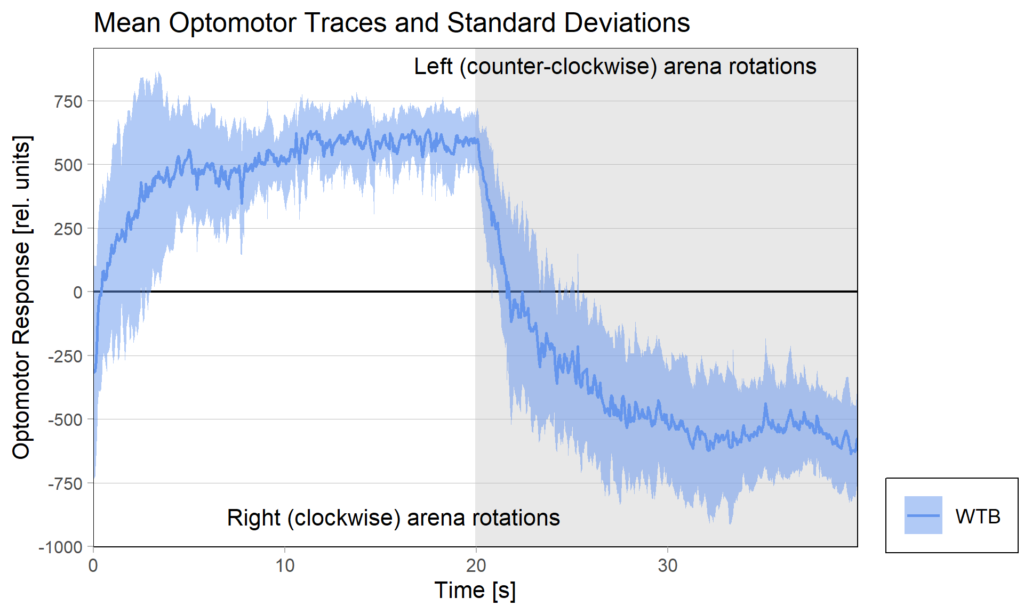
Optomotor traces (right/left) at end of experiment
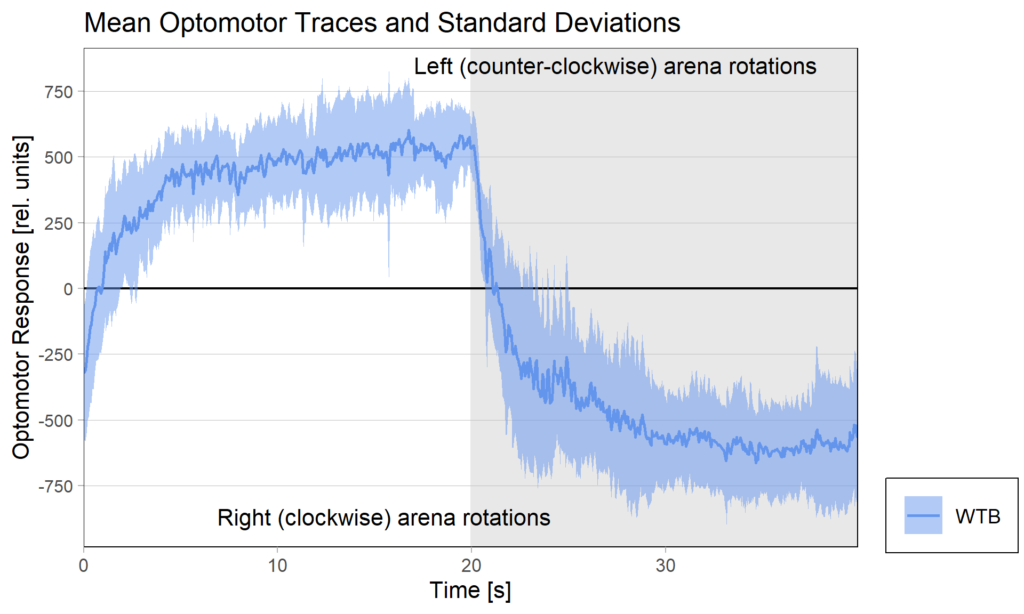
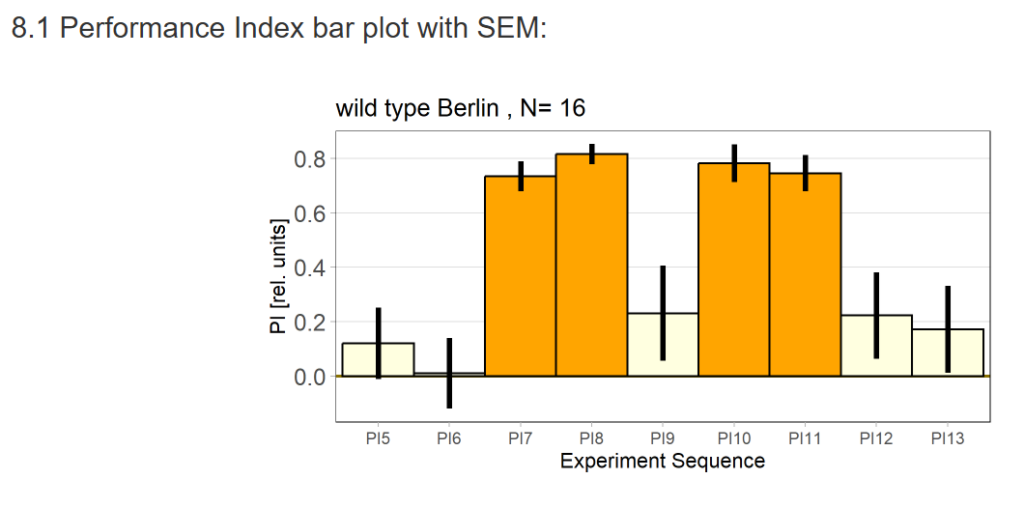
Performance Index bar plot with SEM:
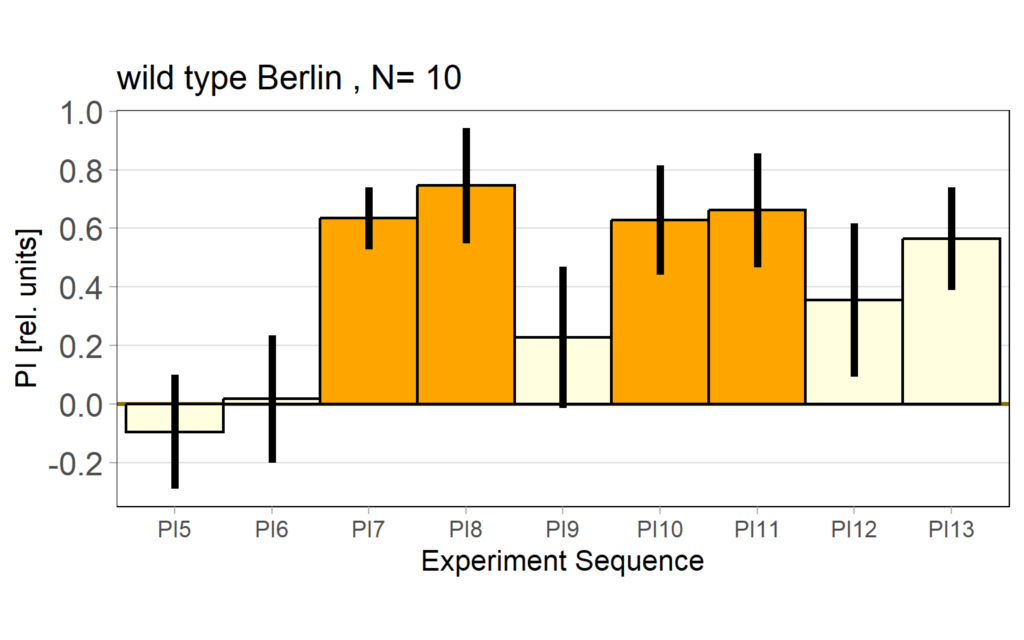
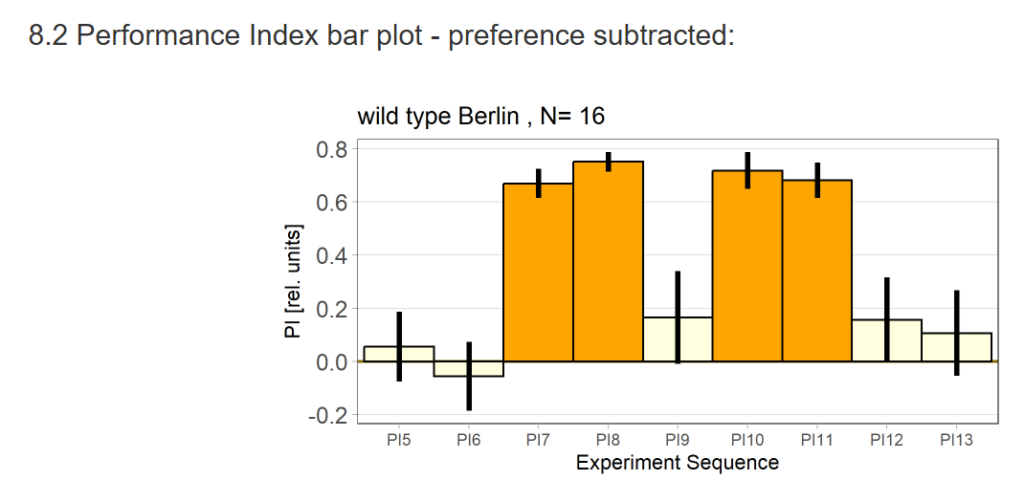
Performance Index bar plot – preference subtracted:
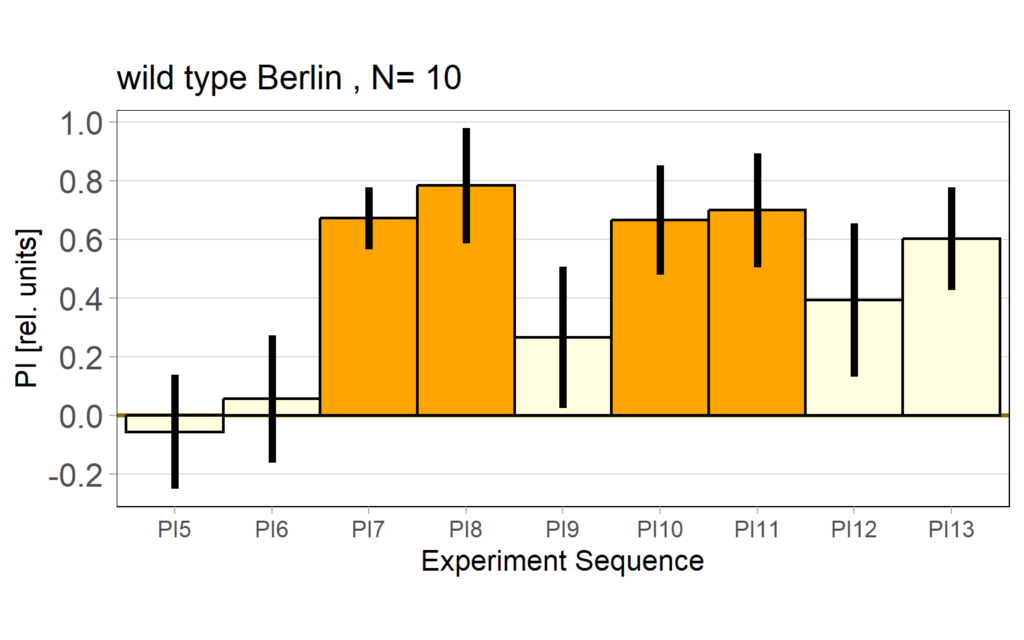
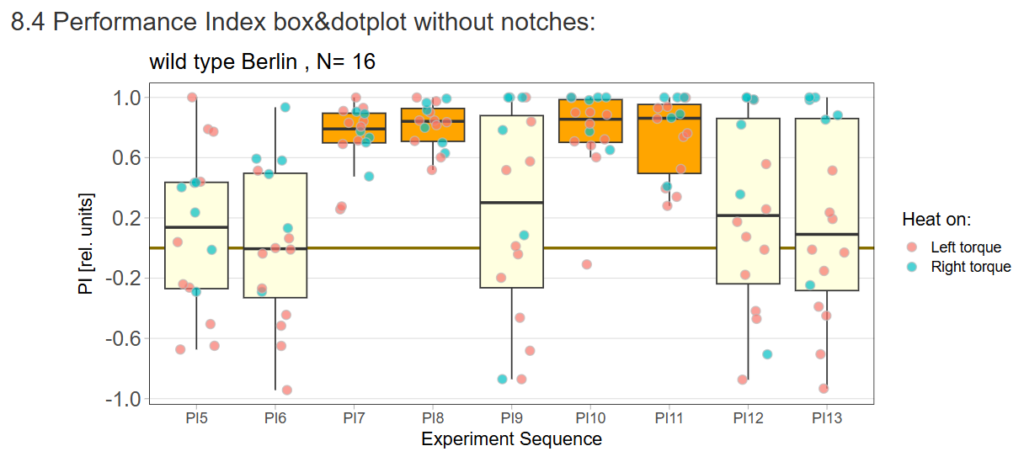
Performance Index box&dotplot without notches:
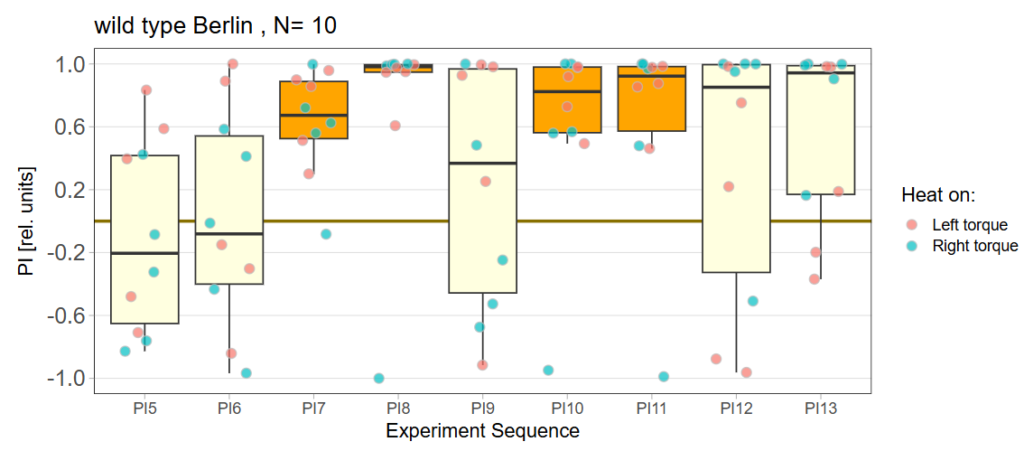
Statistical tests of single groups against zero
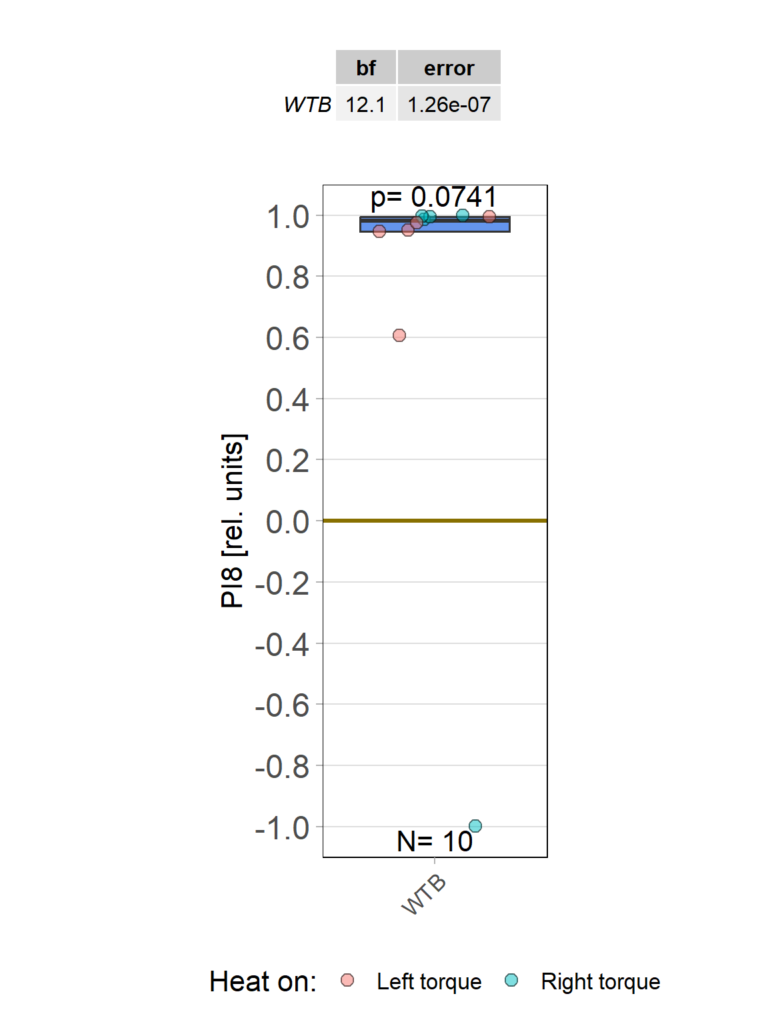
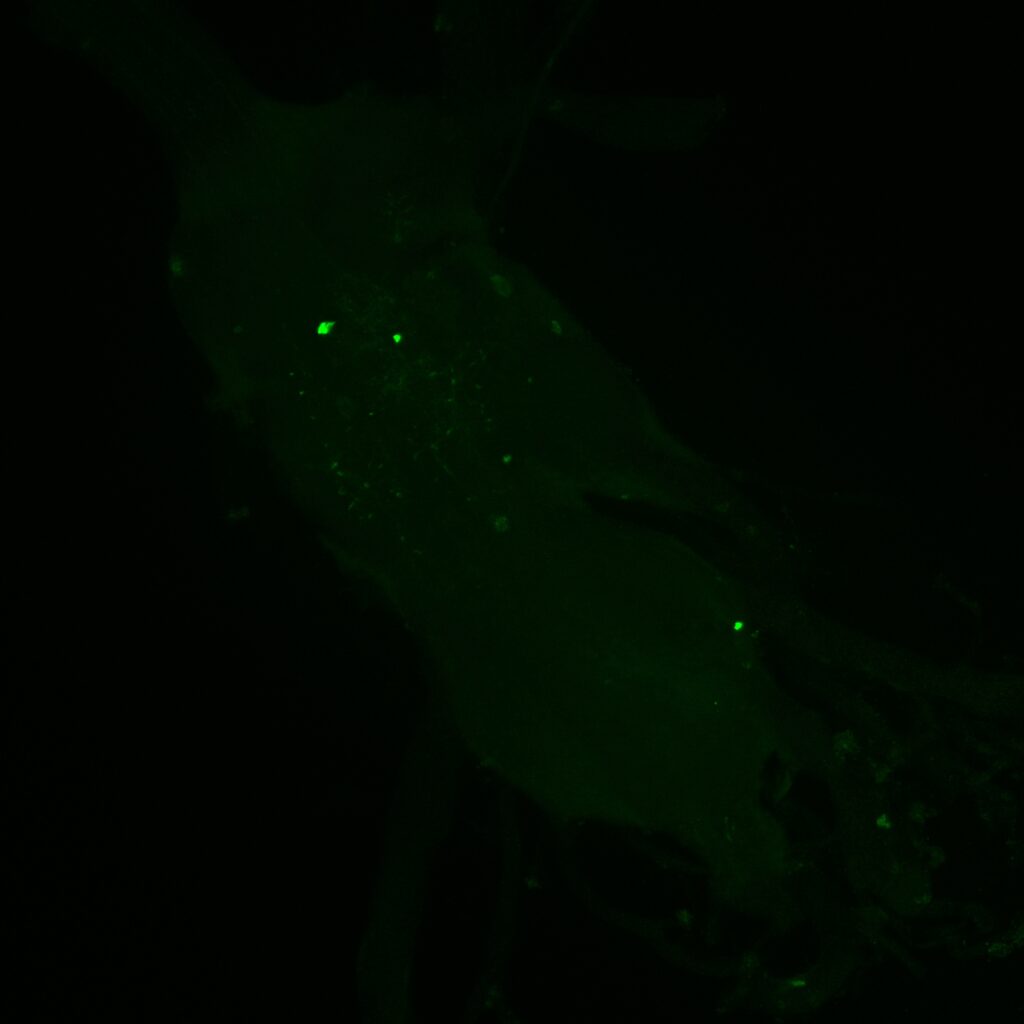
Retro-Tango endogenous GFP Expression
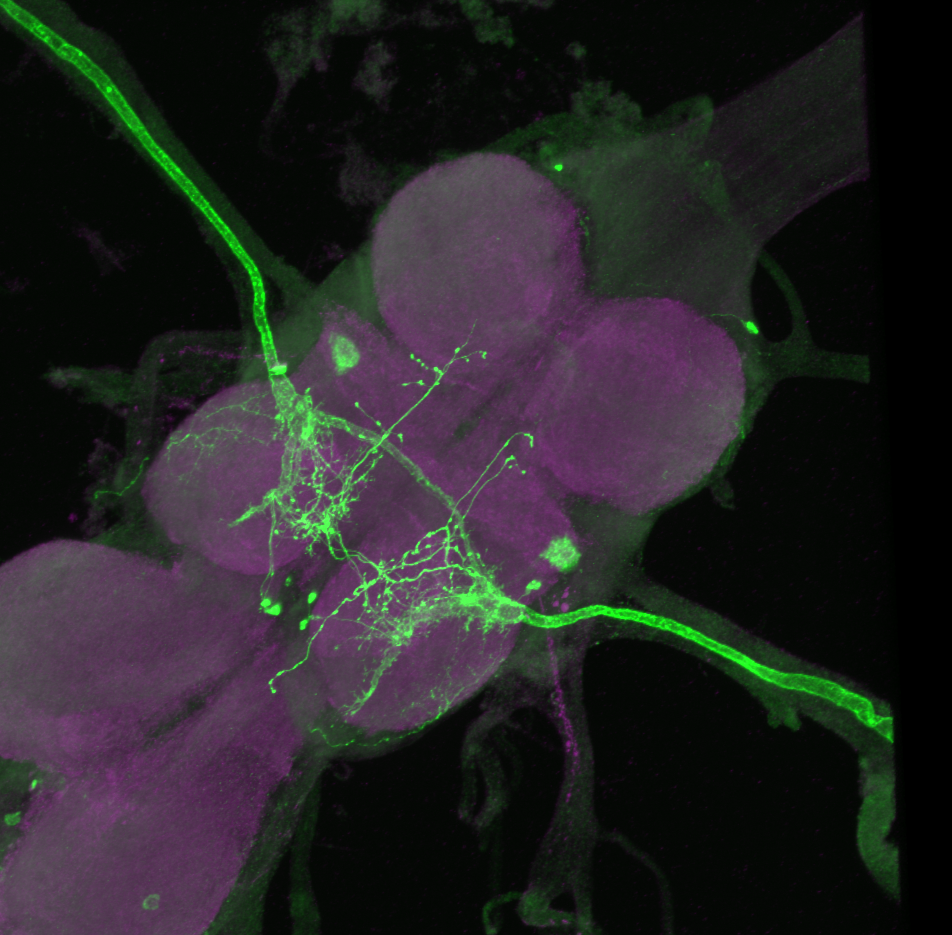
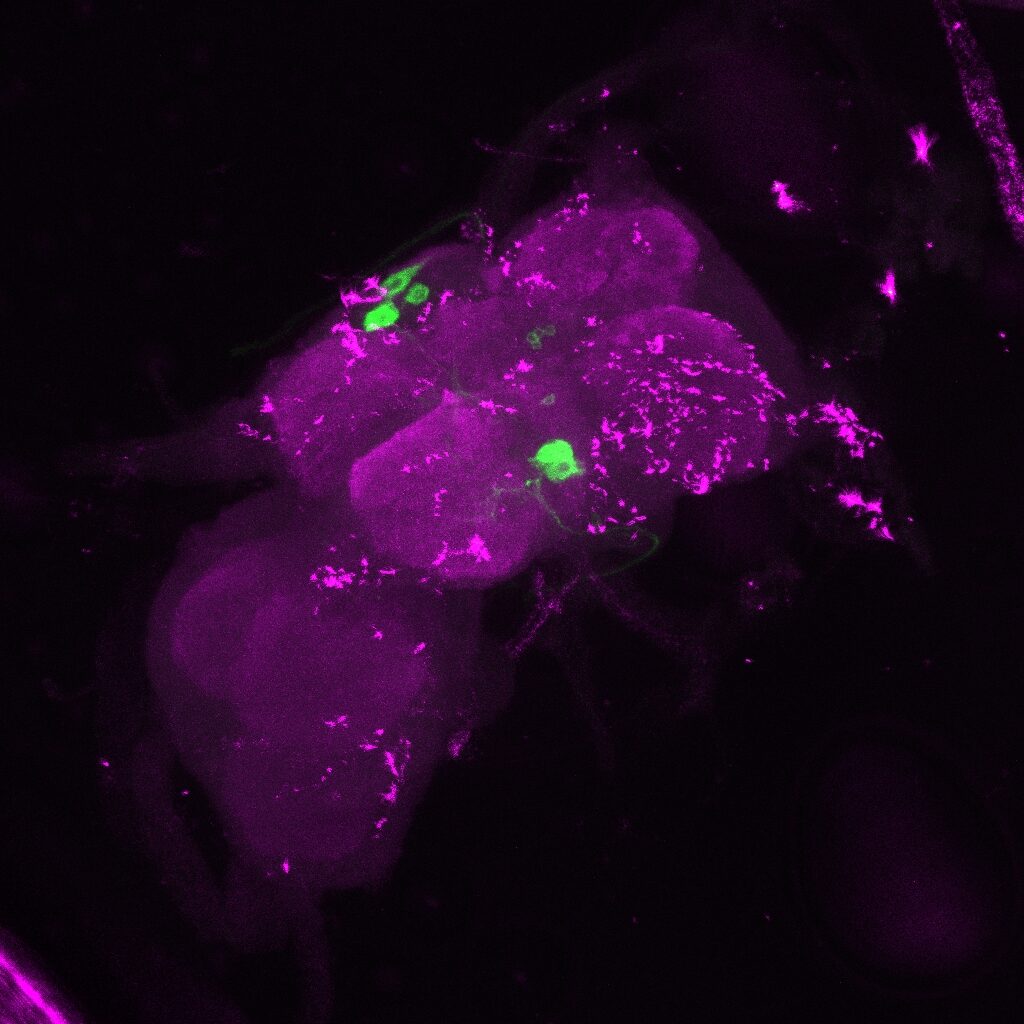
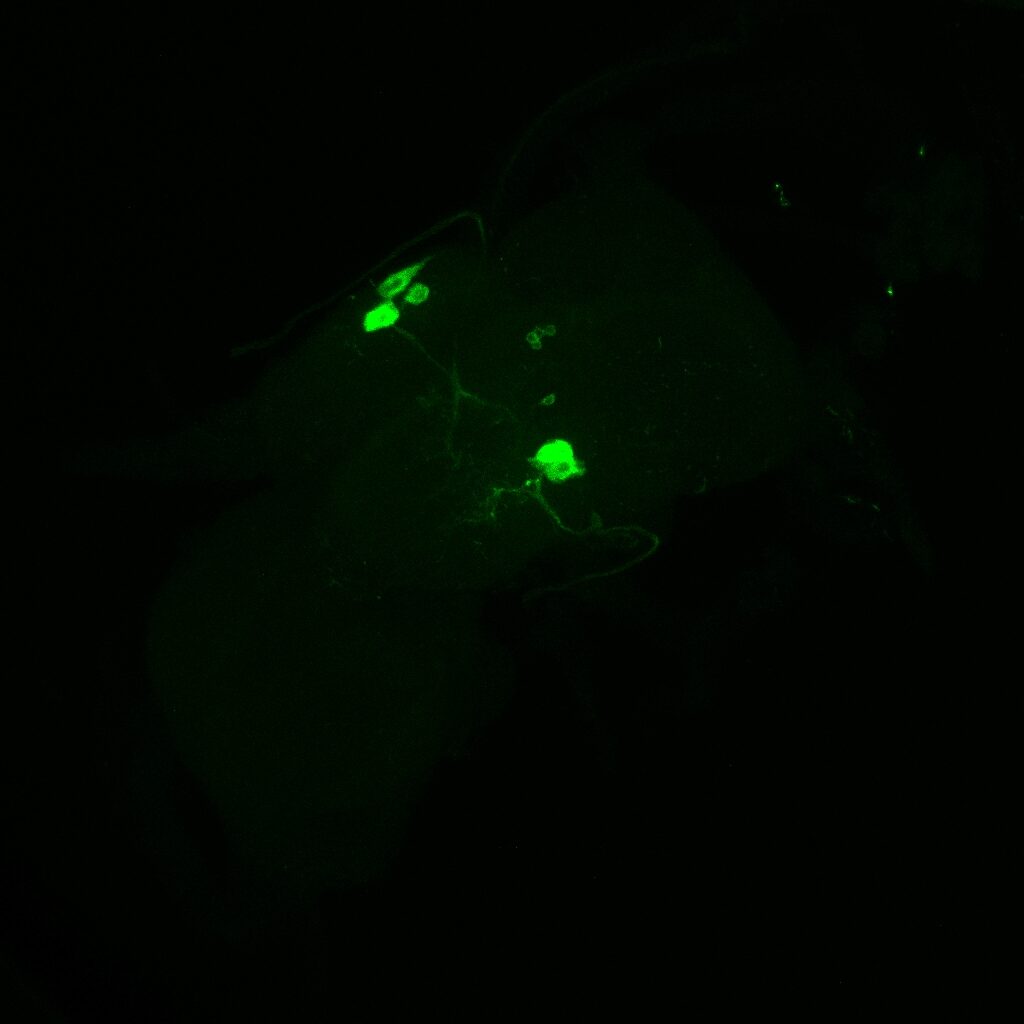
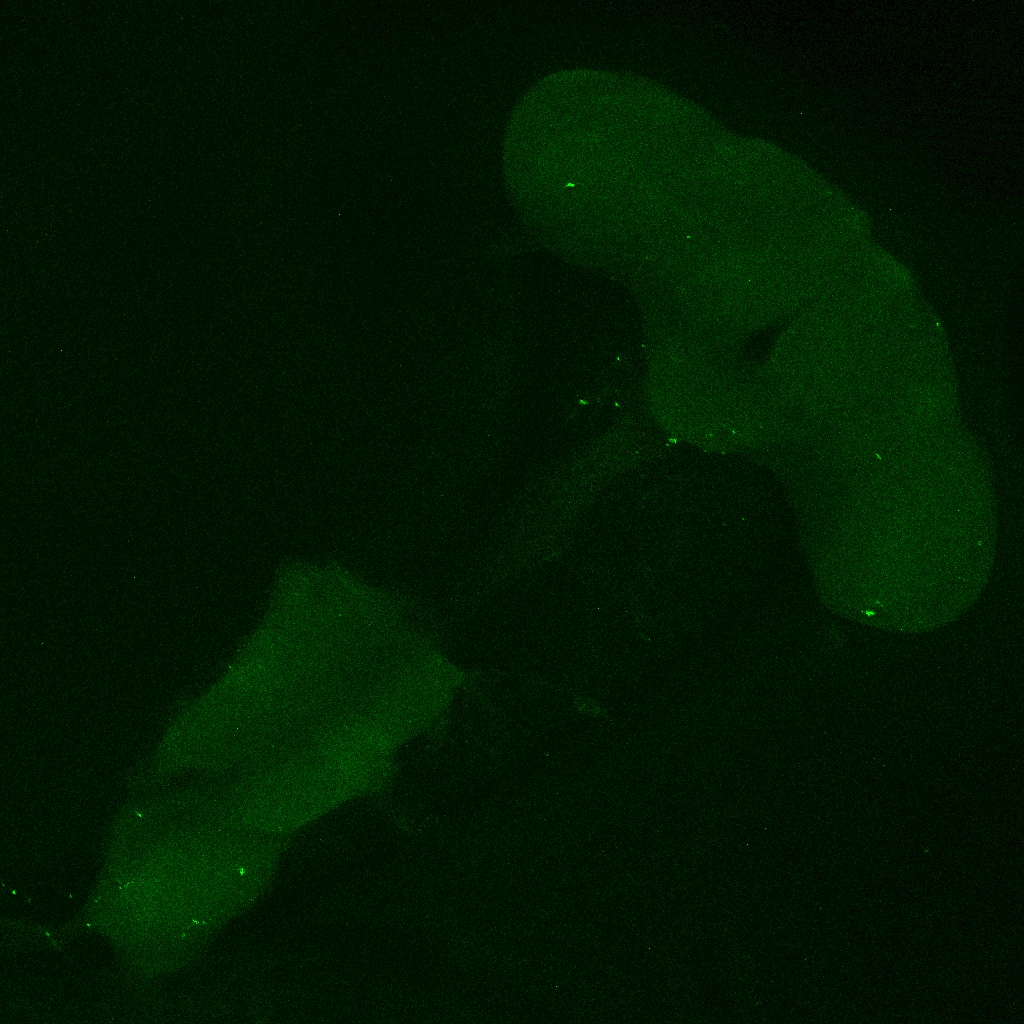
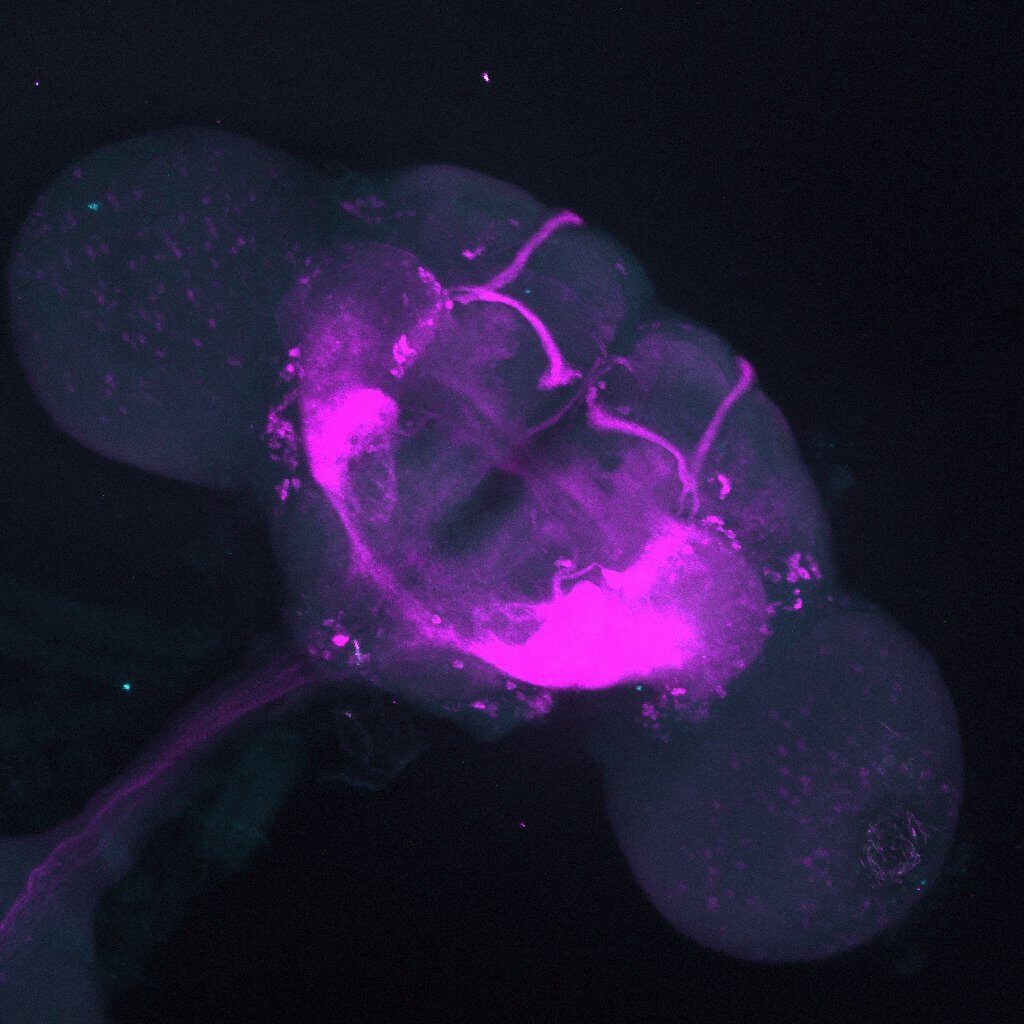
On the right: Endogenous GFP expression (without antibody) from retro tango flies (BDSC: 99661) crossed with GF-Gal4 (BDSC: 79602) driver line.
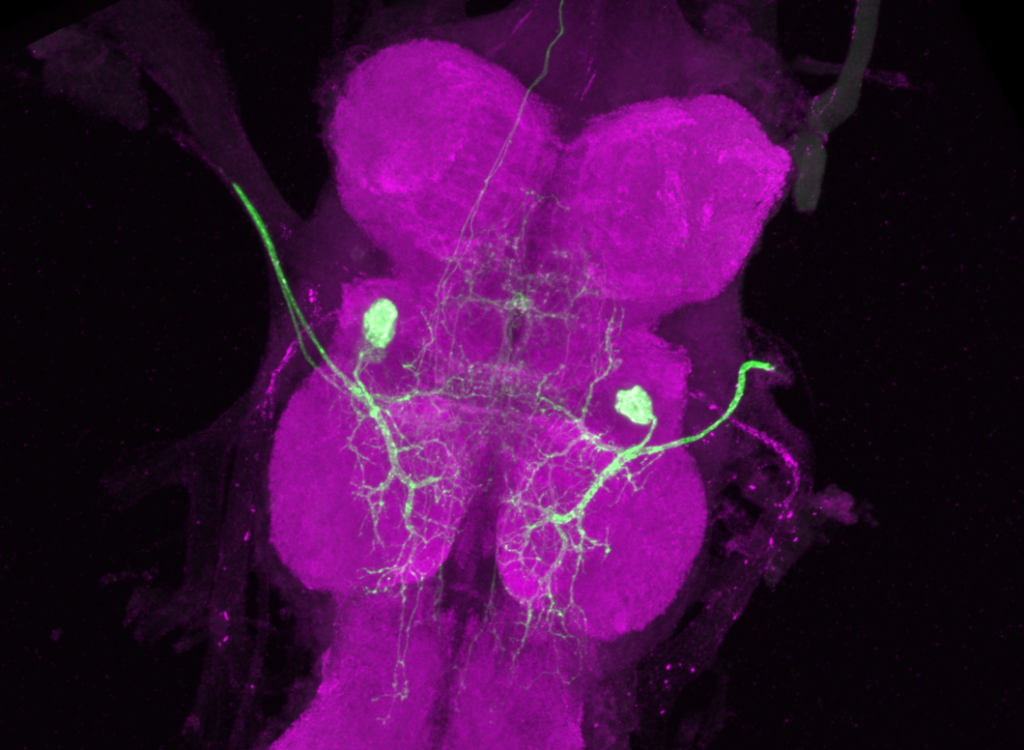
On the left: GFP expression in retro-tango x GF-Gal4 flies from retro-tango publication ( https://doi.org/10.7554/eLife.85041 )
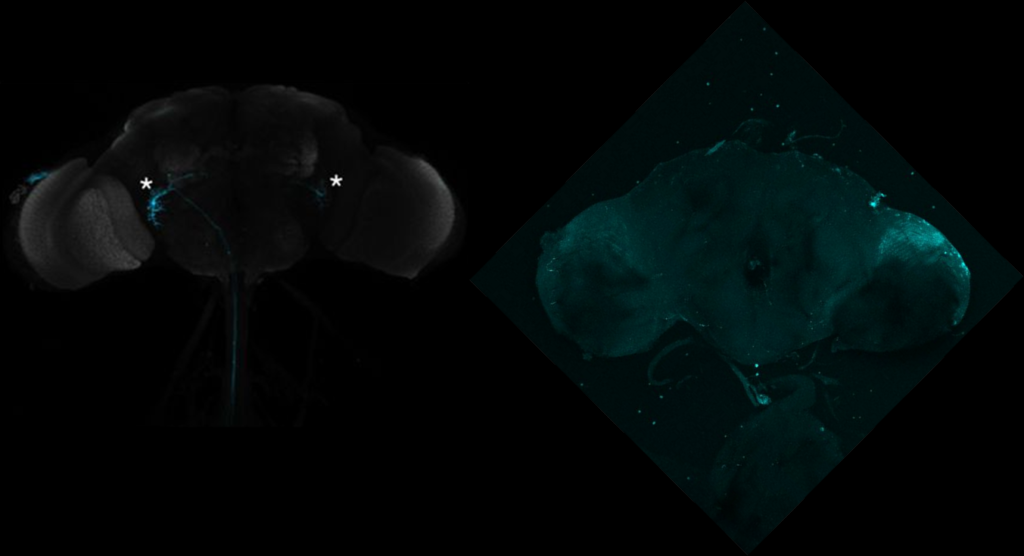
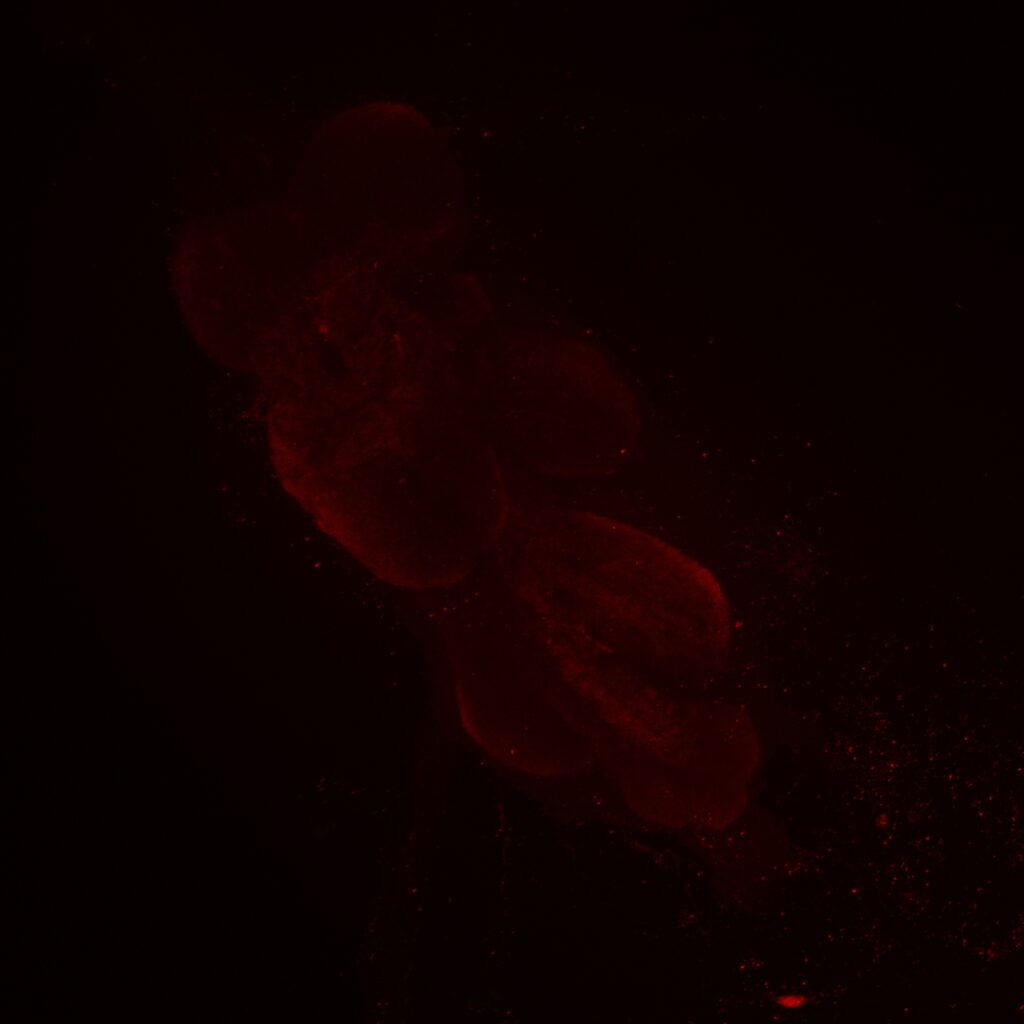
Same image but in retro-tango signal channel:
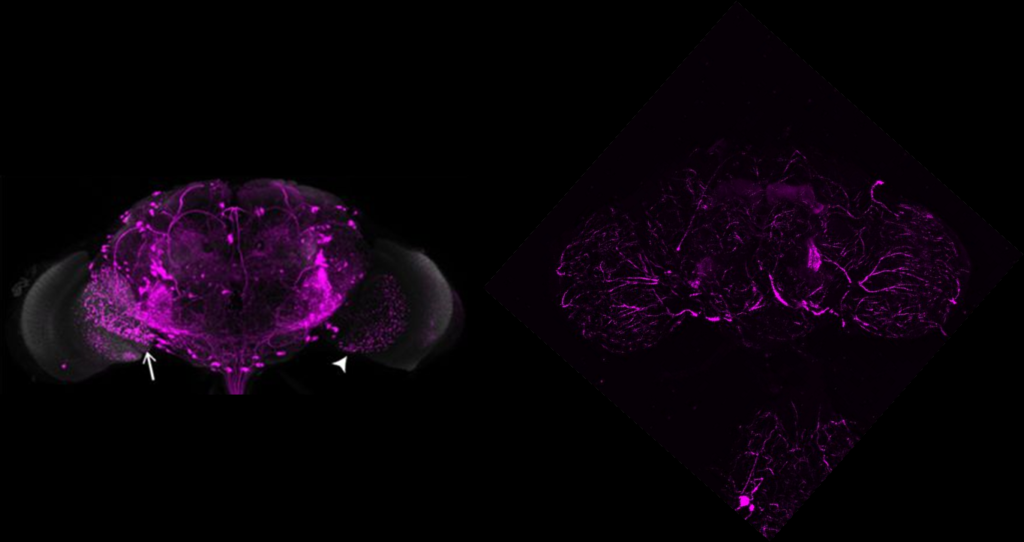
Control Experiments for Driver Line Expression
Giant fiber Gal4 driver (BDSC 79602), green channel shows giant fiber neurons, magenta channel shows background staining with nc82:
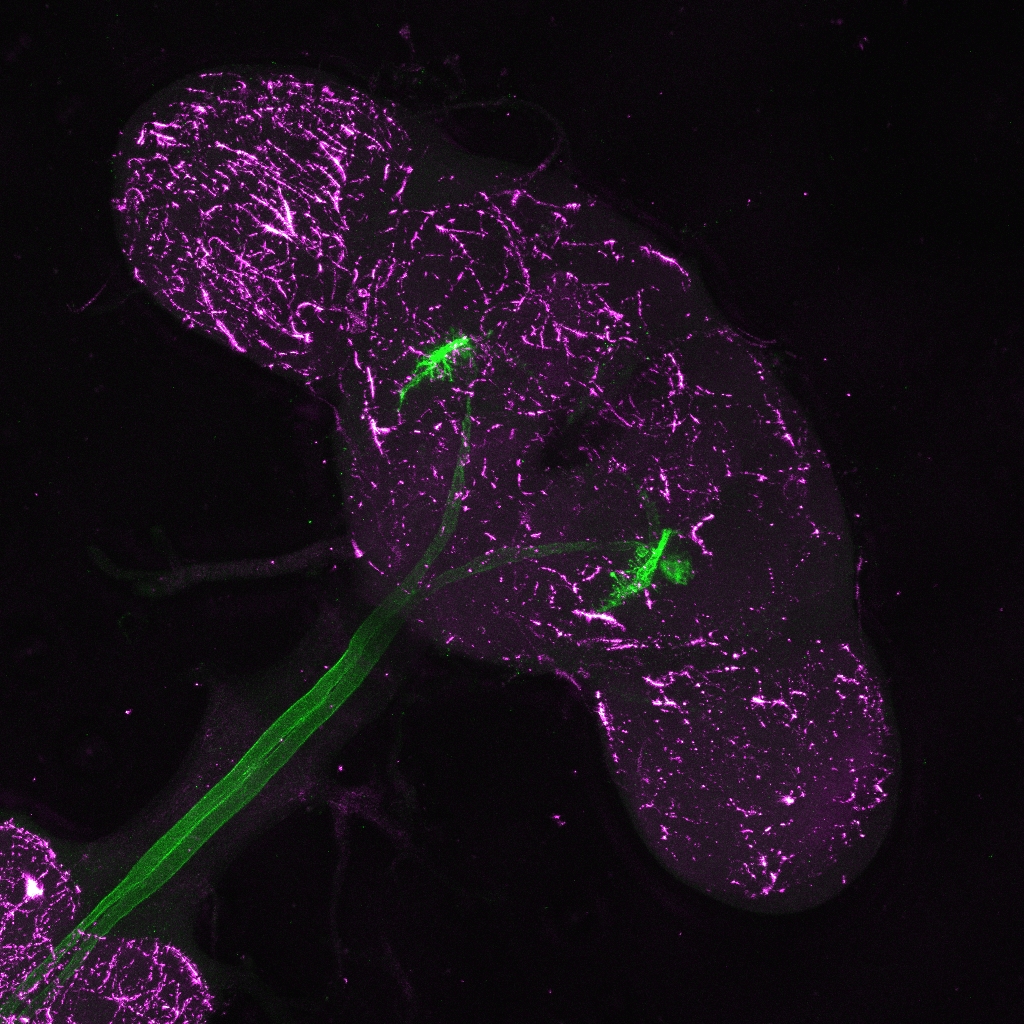
Giant fiber neurons on Codex BANC v626 dataset:
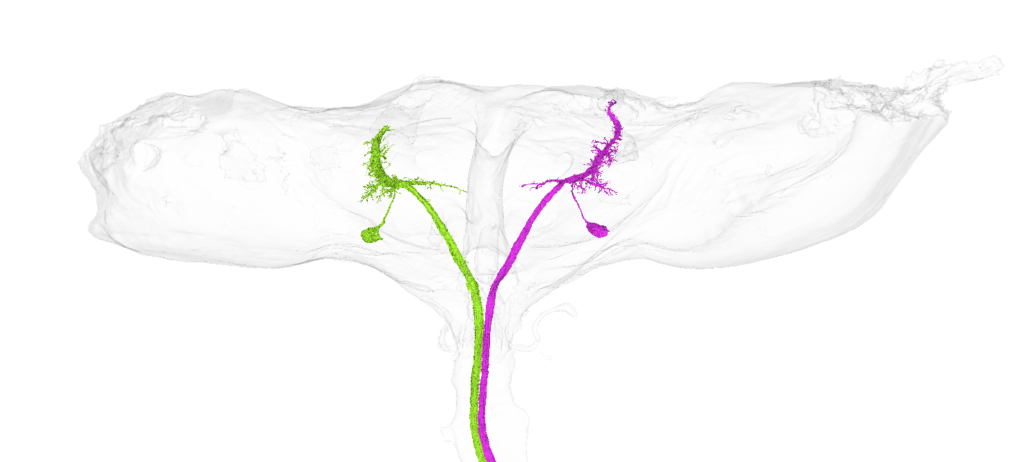
b1 mn driver (SS40980, also targets iii1 mn, iii3 mn):
Janelia image for SS40980:
b3 mn driver (SS48311, also targets hg1 MN, XBI002, WBI009):
Janelia image for SS48311:
b3 mn driver (SS98650):
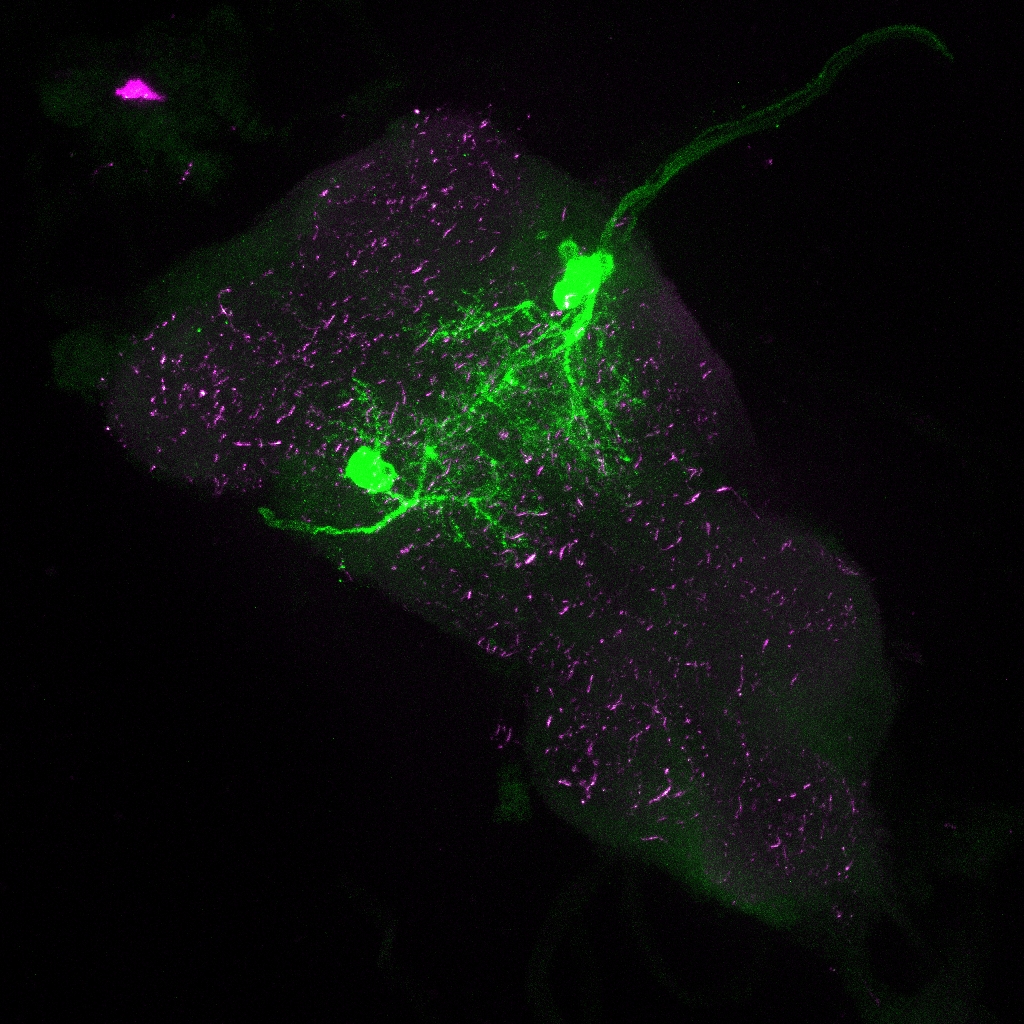
Janelia image for SS98650:
Control Experiments for Driver Line Expression
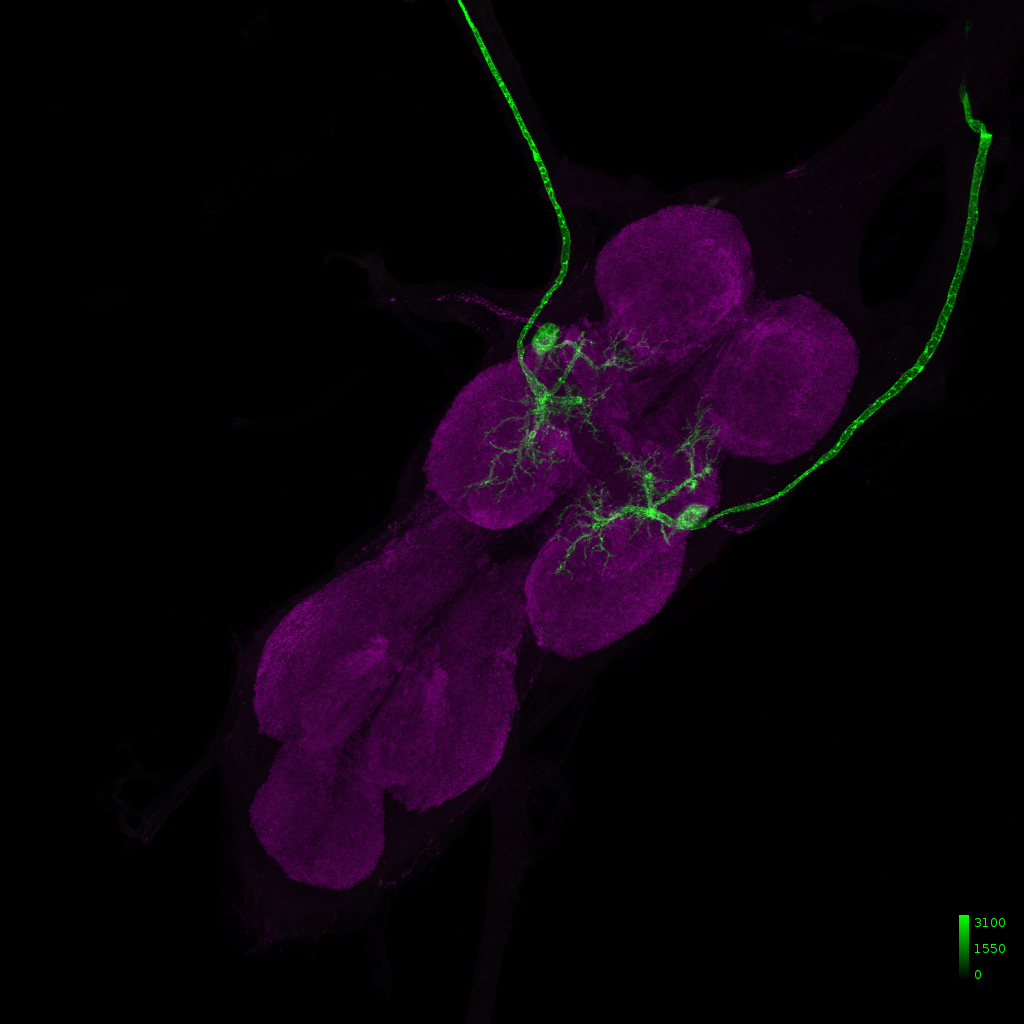
SS45779 – Targeting b3 MN, iii3 MN, hg1 MN
Our line:
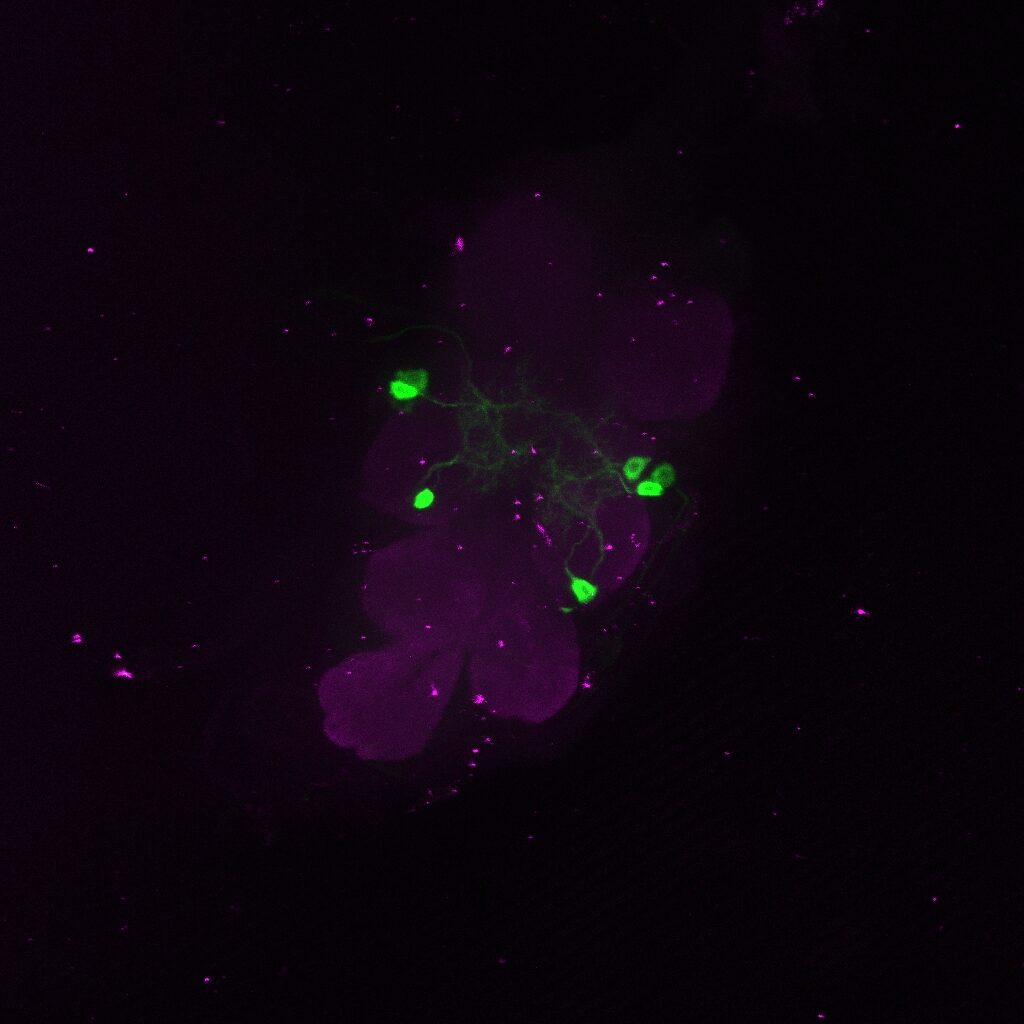
Janelia image:
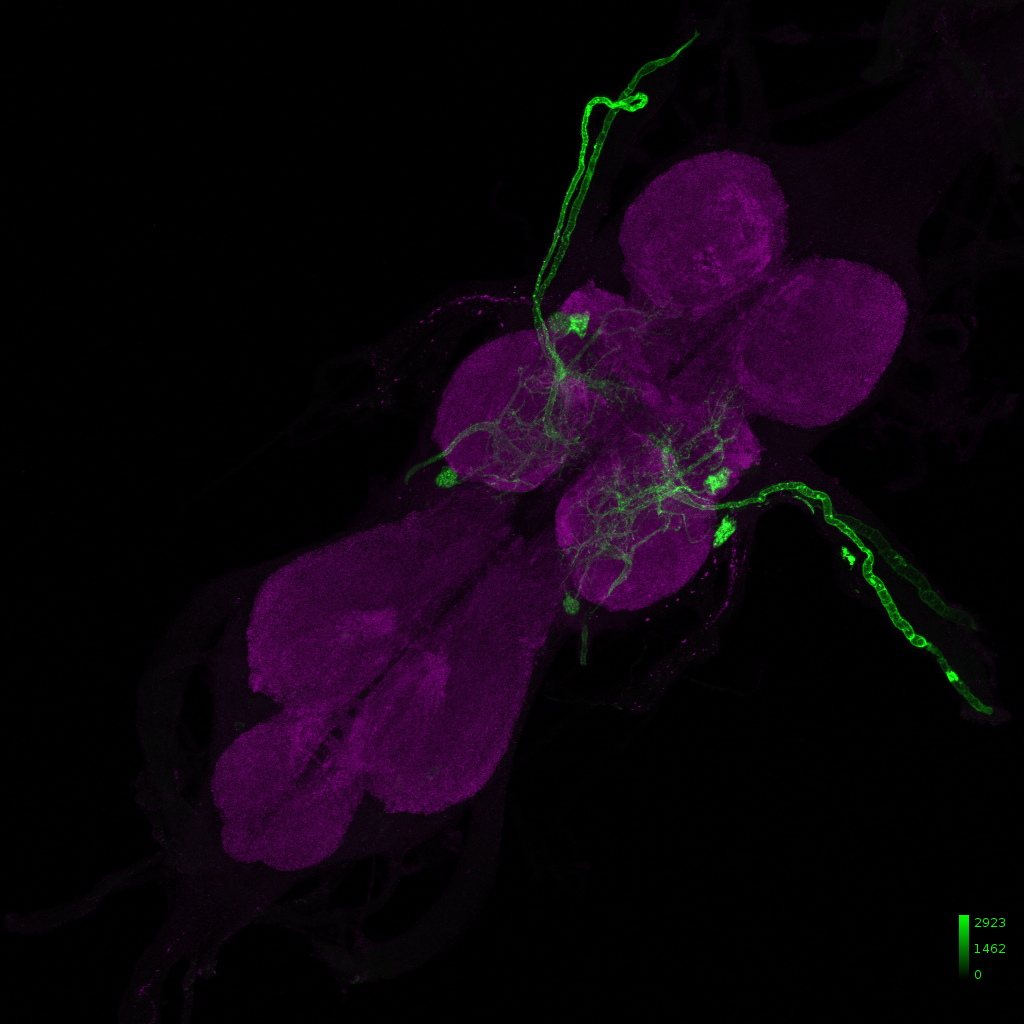
IS69306 – b1 mn driver line
Janelia image:
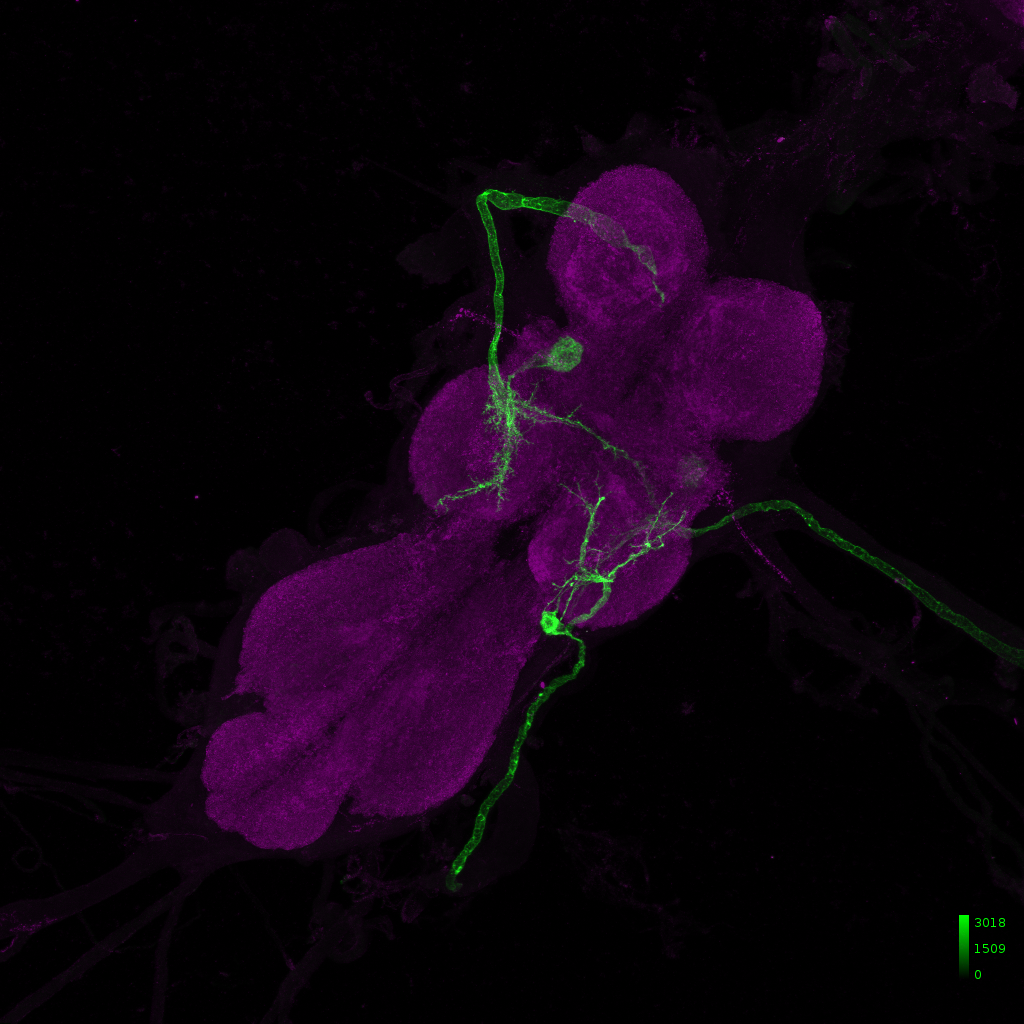
Retro-Tango control crossed with GF-Gal4 (BDSC: 79602), presynaptic neurons in magenta, giant fiber neurons in cyan and background staining in gray
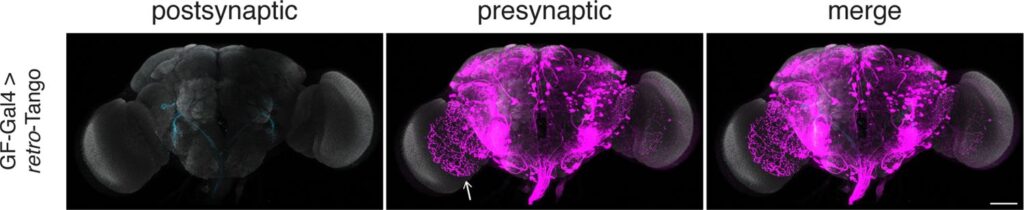
Retro-Tango x GF-Gal4 from Sorkac et. al (2023)
rut and rad expression in b1 and b3 motor neurons, I looked for cells express 6 genes (rut, rad, FoxP, VGlut, AD TF, DBD TF) at the same time above certain threshold:
| Driver Line | AD TF | DBD TF | Targeted cells | Cells express all 6 genes at more than 0 | Cells express all 6 genes at more than 1 fold | Cells express all 6 genes at more than 2 fol |
| SS40980 | CG12680 | Cep104 (CG10137) | b1 MN, iii1 MN, iii3 MN | 0 | 0 | 0 |
| SS45779 | Ubx | ab | b3 MN, iii3 MN, hg1 MN | 158 | 0 | 0 |
| SS47160 | Ubx | hry | b2 MN, b3 MN, iii4 MN, hg1 MN, hg3 MN | 1 | 0 | 0 |
| SS48311 | Ubx | NetB | b3 MN, hg1 MN, XBI002, WBI009 | 78 | 44 | 2 |
| SS49806 | ??? | tim | b3 MN, hg1 MN, HBI017 | – | – | – |
| SS52405 | Ubx | CG4168 | b3 MN, i2 MN, hg3 MN | 46 | 25 | 1 |
| IS69306 | CG14388 (CG31345) | Awh | b1 driver | 0 | 0 | 0 |
| SS98650 | ct | Ubx | b3 driver | 76 | 39 | 0 |
| IS38497 | beat-IIIb | bab1 | b1&b3 | 34 | 15 | 1 |
Retro-tango confocal images and Torquemeter practice results
Retro-tango:
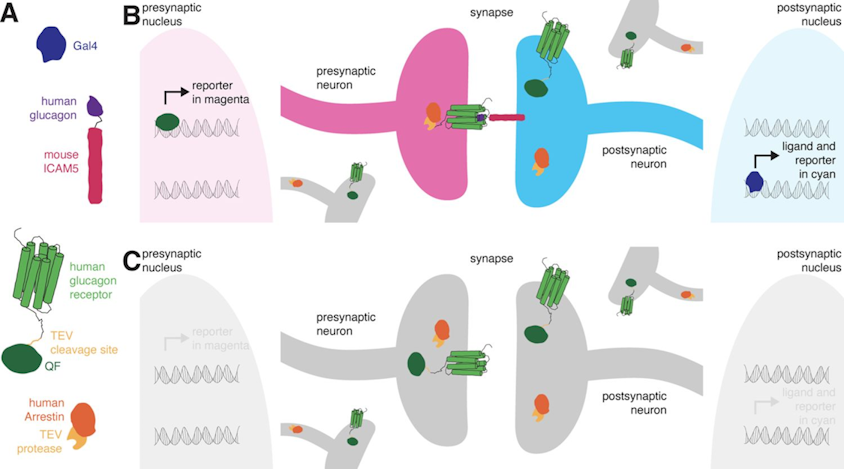
(2023) retro-Tango enables versatile retrograde circuit tracing in Drosophila eLife 12:e85041. https://doi.org/10.7554/eLife.85041
Retro-tango genotype: y[1] w[*] P{y[+t7.7] w[+mC]=QUAS-mtdTomato-3xHA.S}su(Hw)attP8; P{y[+t7.7] w[+mC]=retro-Tango(panneuronal)}attP40/SM6b; P{y[+t7.7] w[+mC]=10xUAS-retro-Tango(ligand)-P2A-EGFP-F}attP2
Crossed with IS69306 (BDSC 601295 and 75552) split-Gal4 driver to express in b1 motor neurons. CNSs stained with anti-GFP Rabbit, (2nd ab: goat anti-rabbit 488), anti-HA Rat (2nd ab: goat anti-rat 555), nc82 (2nd ab: goat anti-mouse 647)
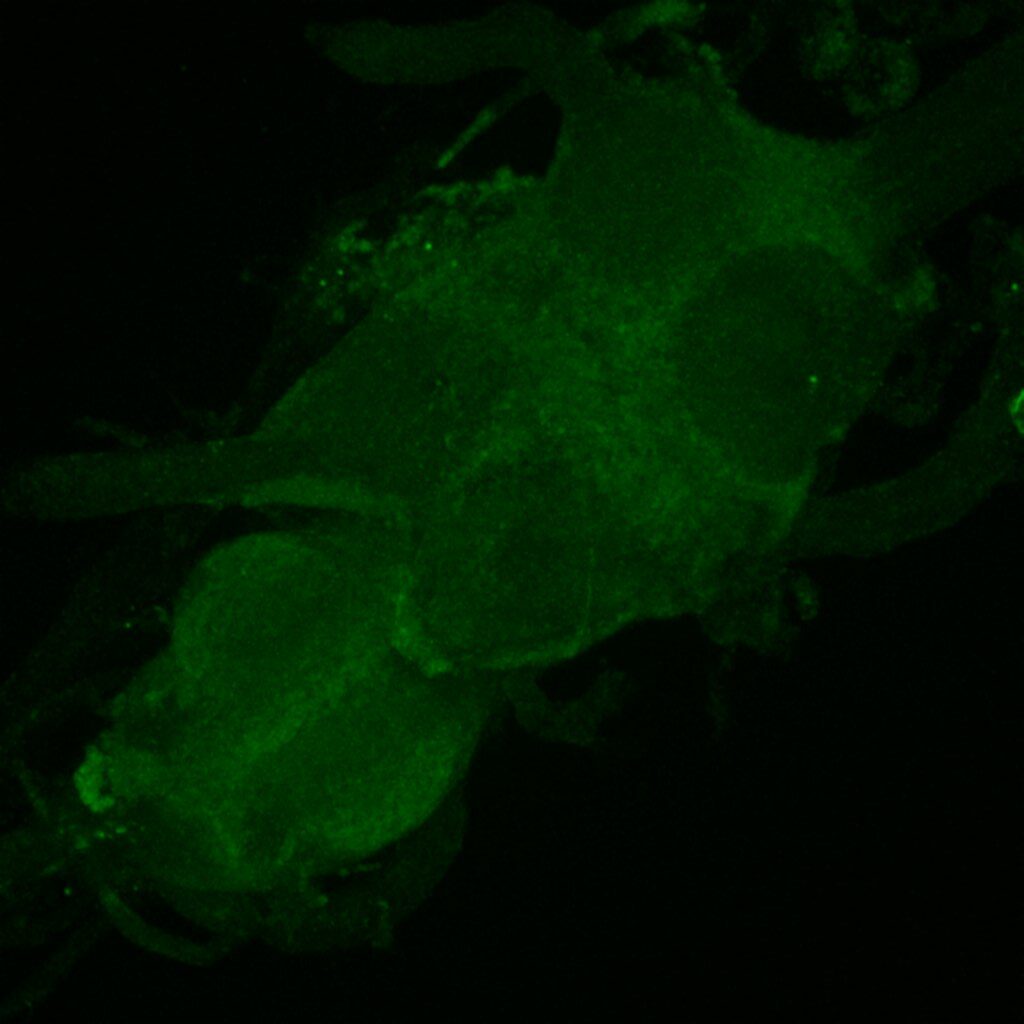
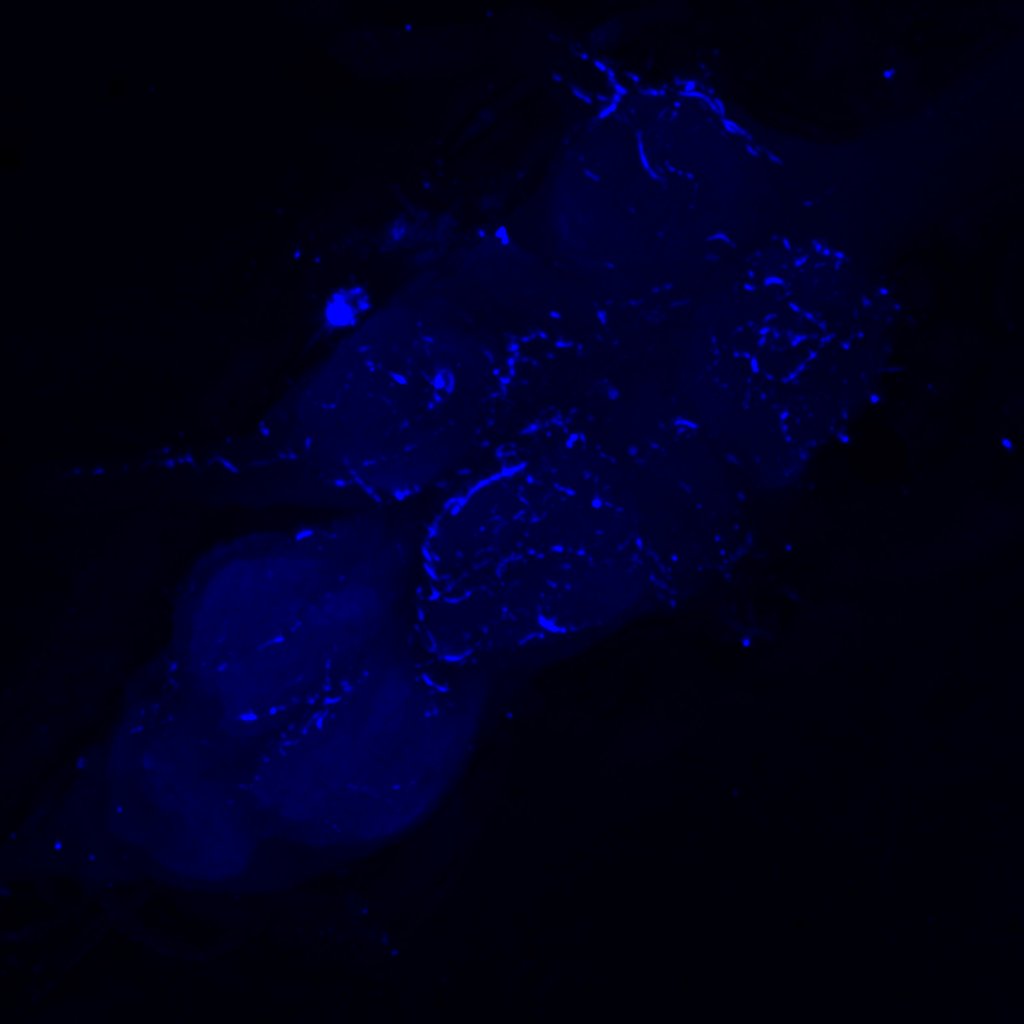
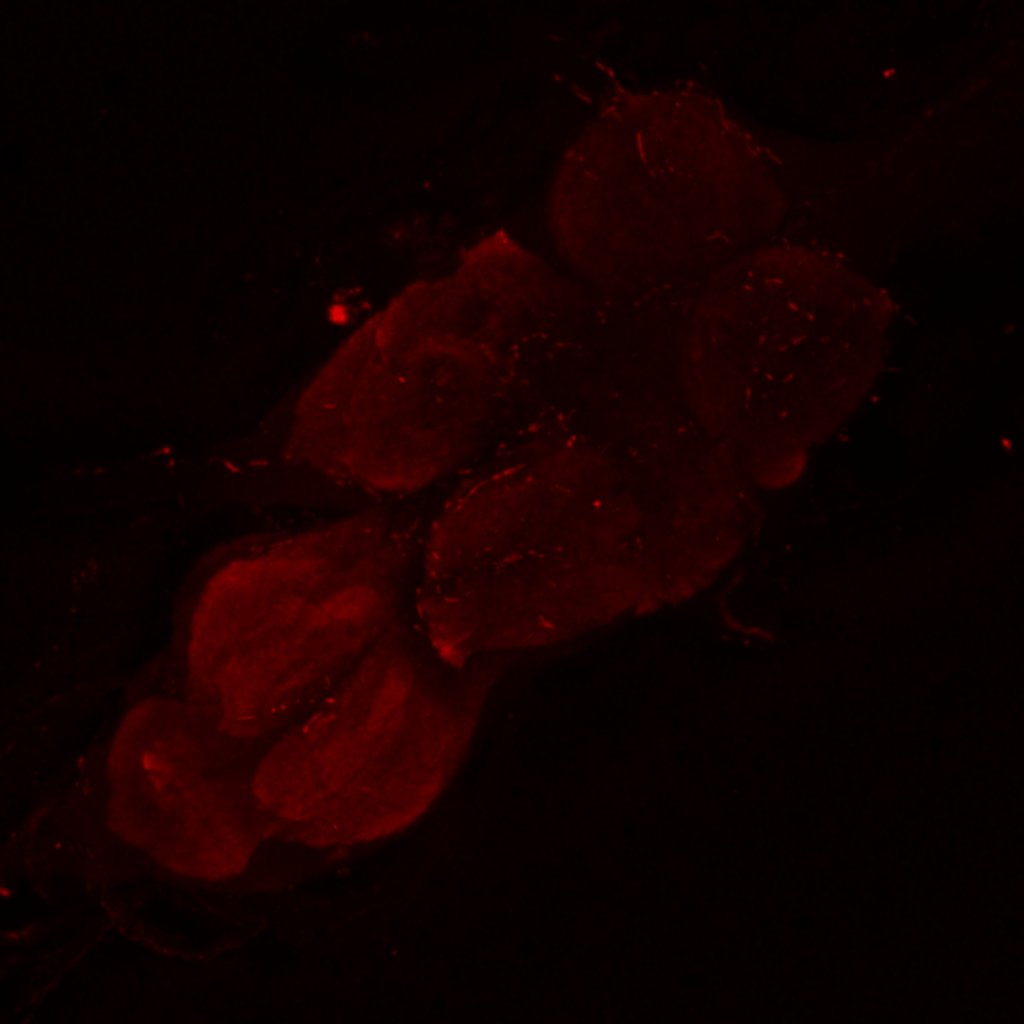
Channel 1 (Green: Laser Line ( 496 nm) Intensity: 23.99%) Spectral Positions/Gain/Offset: (501nm – 556nm) / 865.7 / -0.03
Channel 2 (Blue: Laser Line ( 561 nm) Intensity: 13.10%) Spectral Positions/Gain/Offset: (566nm – 639nm) / 537.7 / 0.03
Channel 3 (Red: Laser Line ( 633 nm) Intensity: 23.99%) Spectral Positions/Gain/Offset: (644nm – 776nm) / 802.9 / -0.01
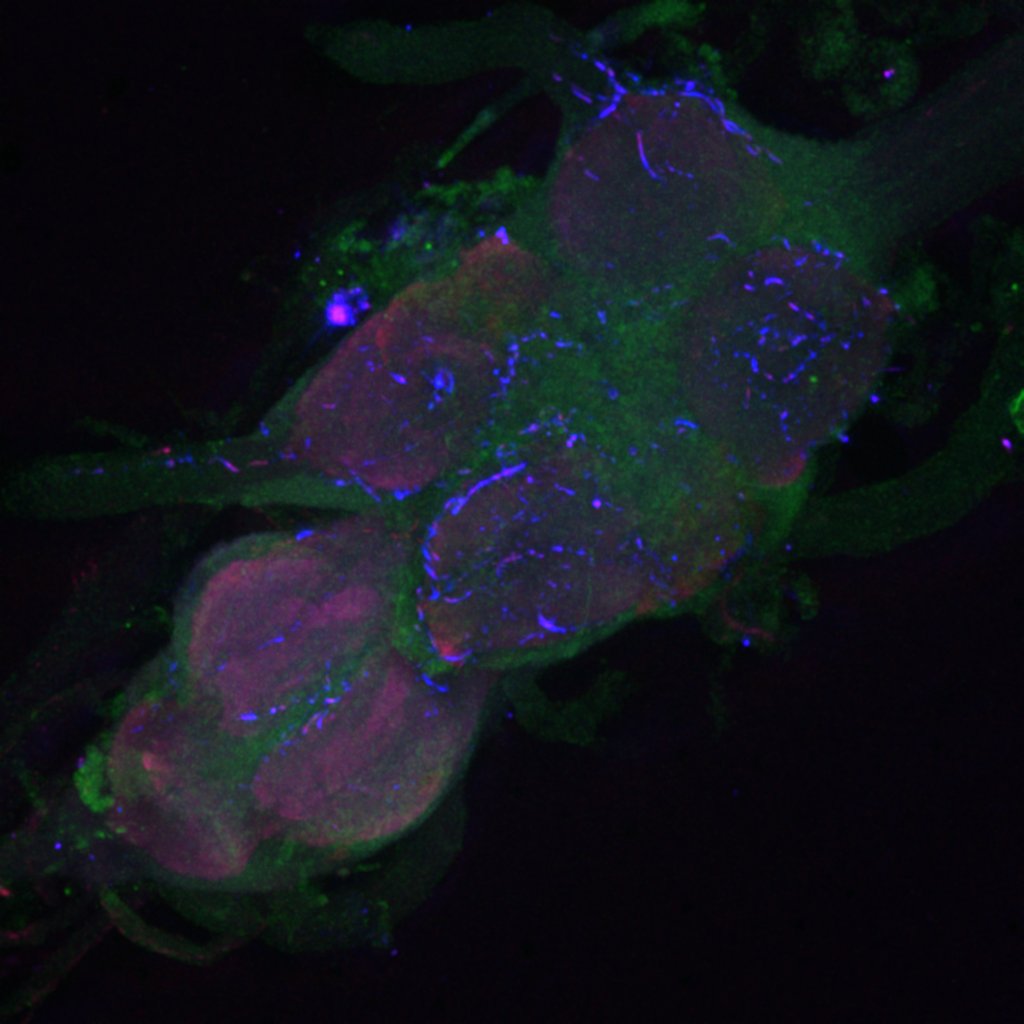
Second line retro-tango crossed with SS98650 (split-Gal4 driver line from Janelia targeting b3 motor neurons)
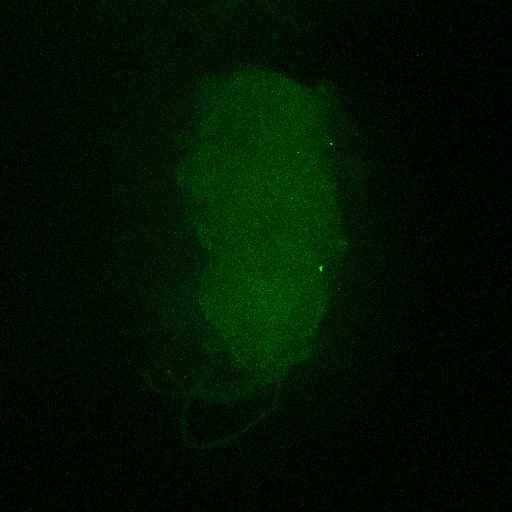
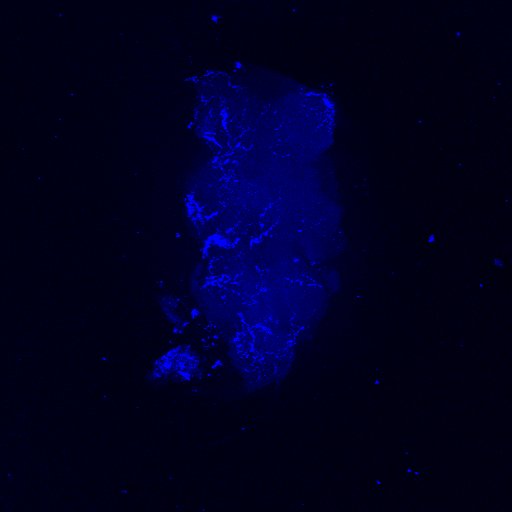
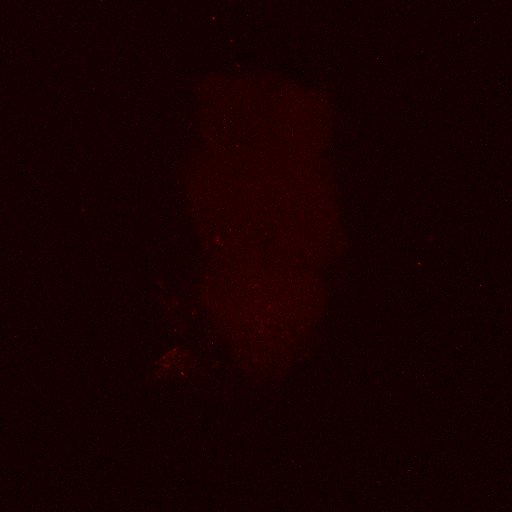
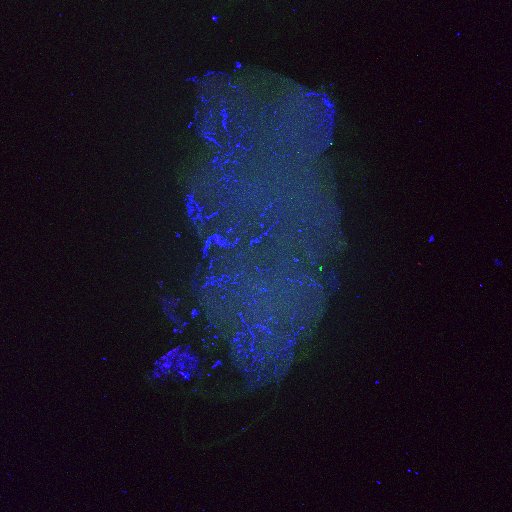
Settings are same as above
Torquemeter Practice with WTB Flies
N=10 out of 24 glued flies
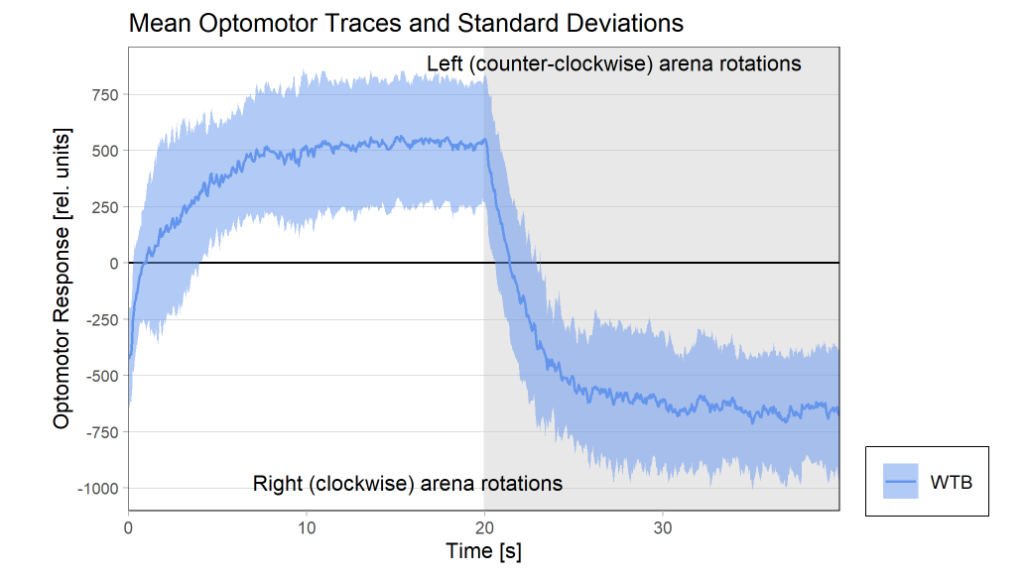
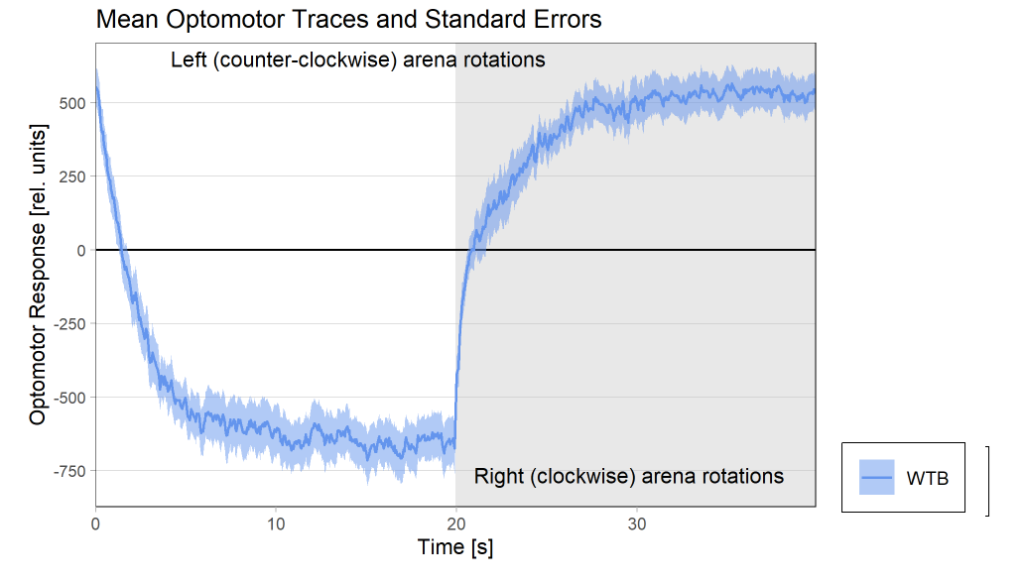
Optomotor at start:
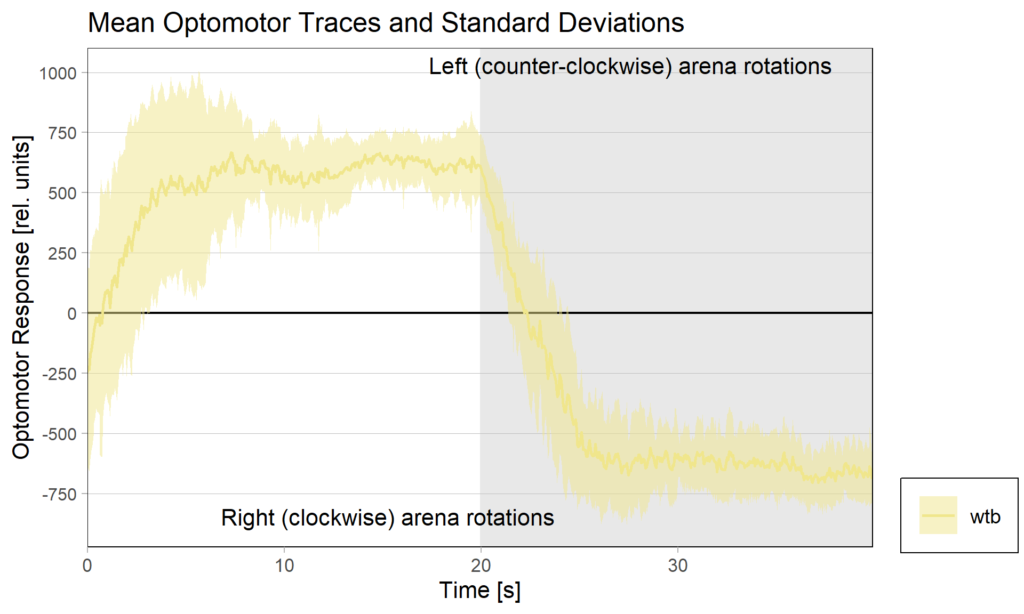
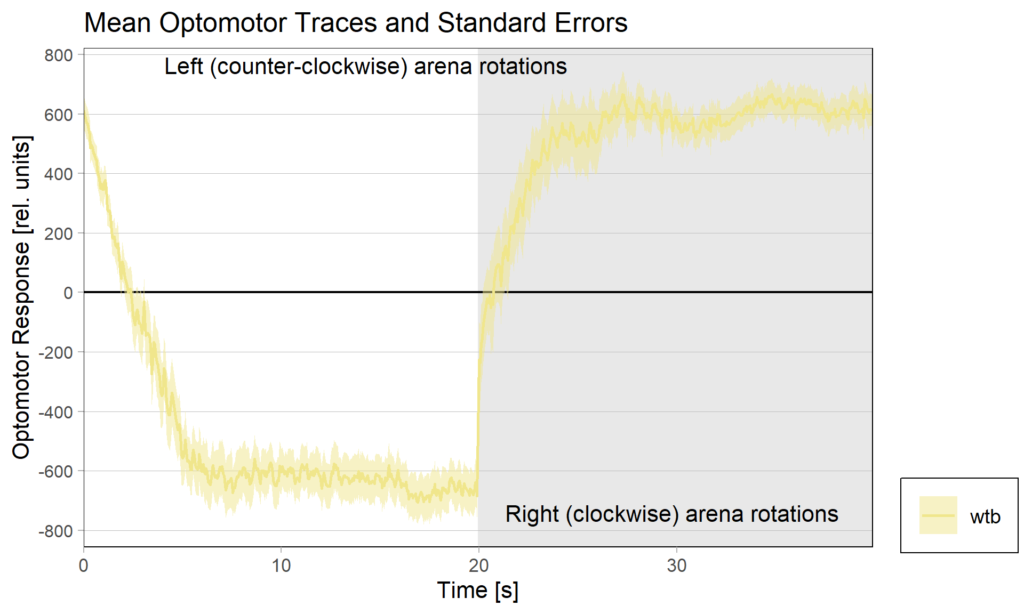
Optomotor end:
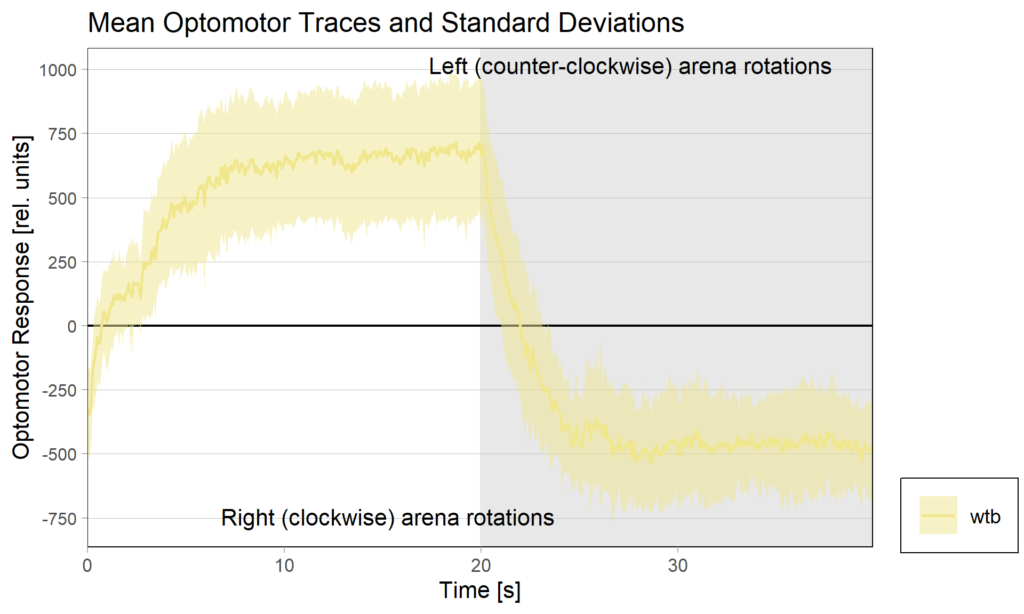
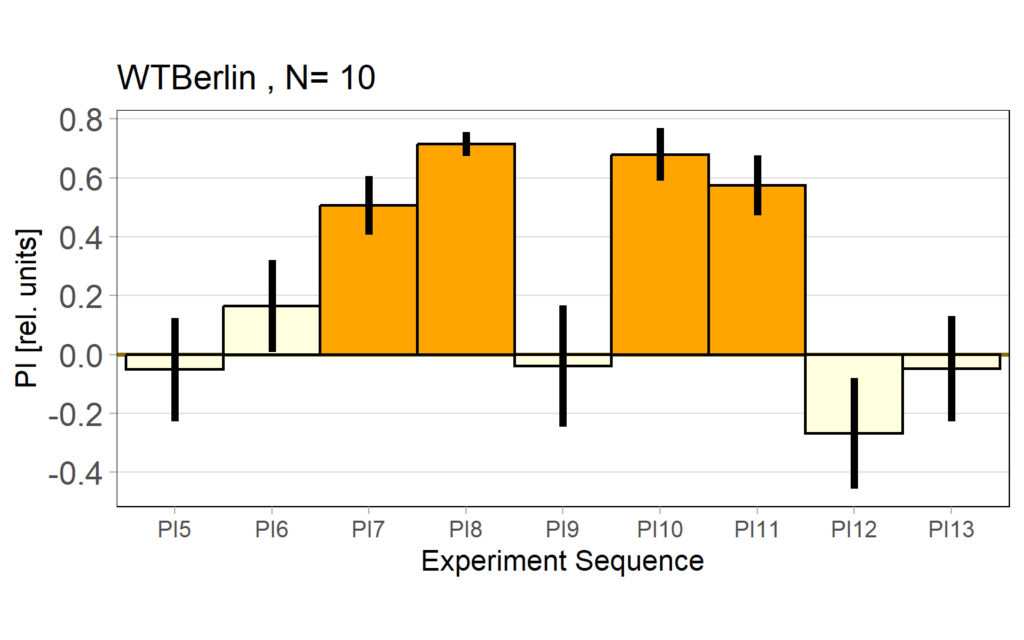
Performance index:
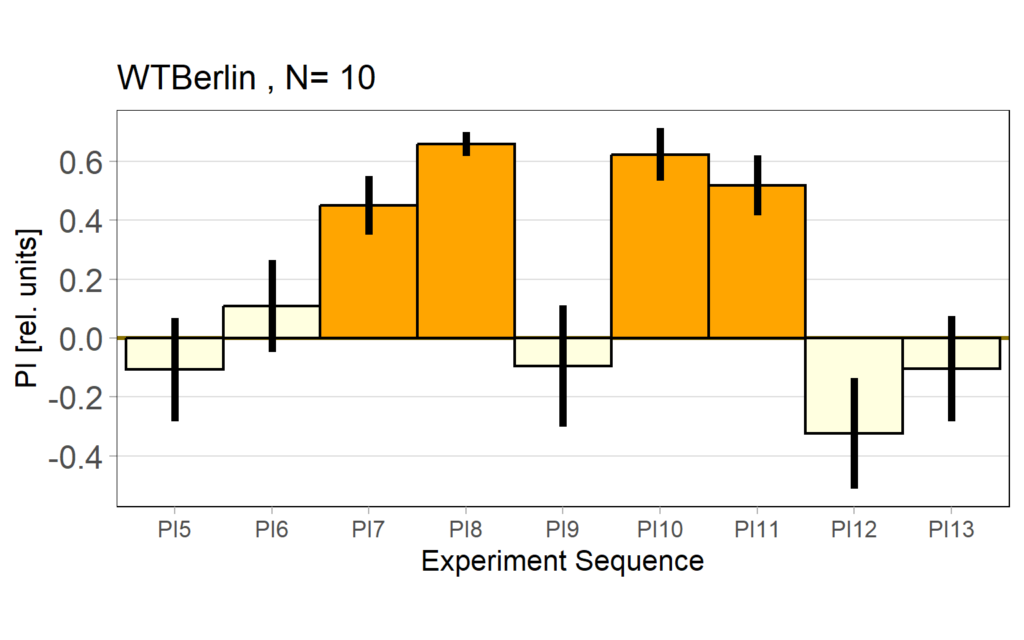
Performance subtracted:
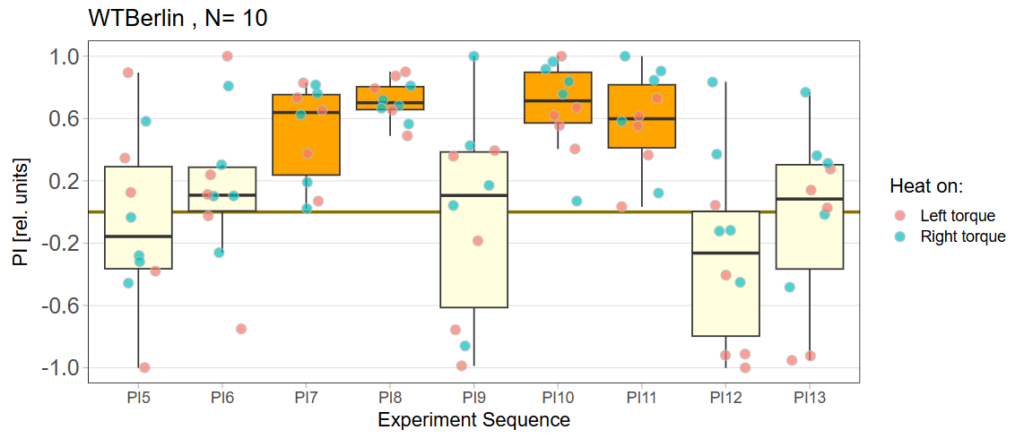
rut and rad expression in ventral nerve cord:
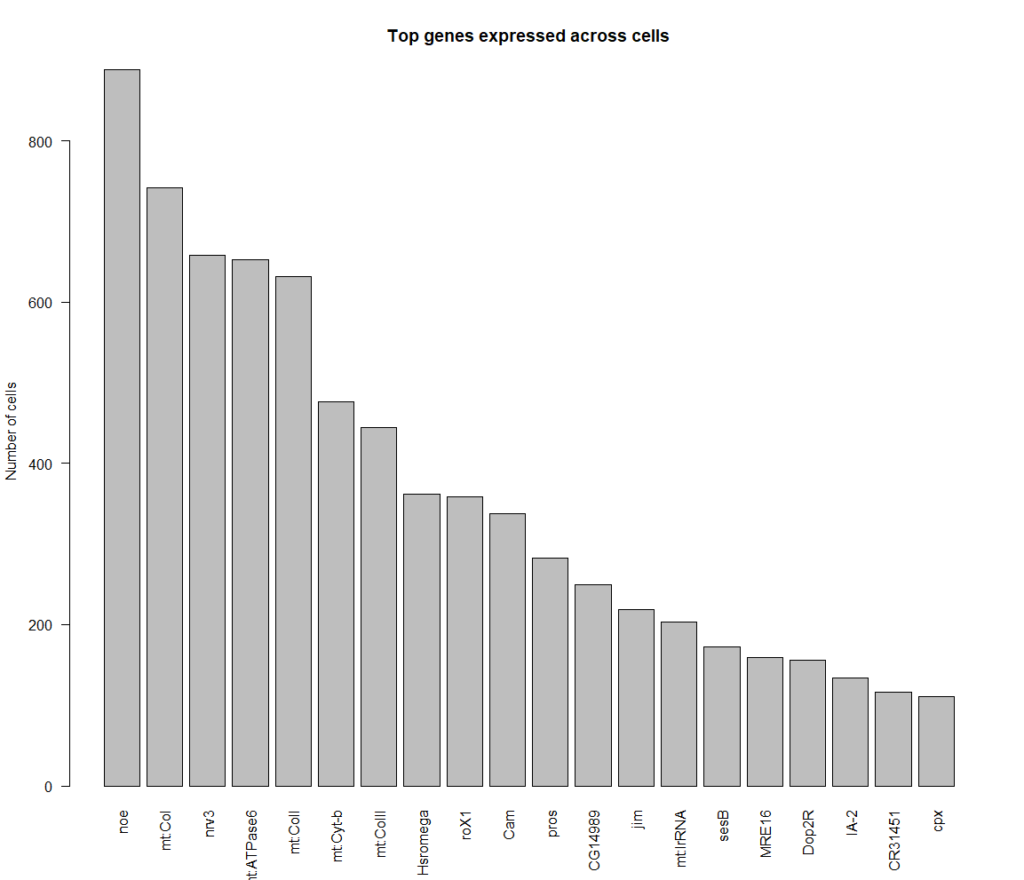
cells that express all 4 genes more than 2 fold, 889 cells in total
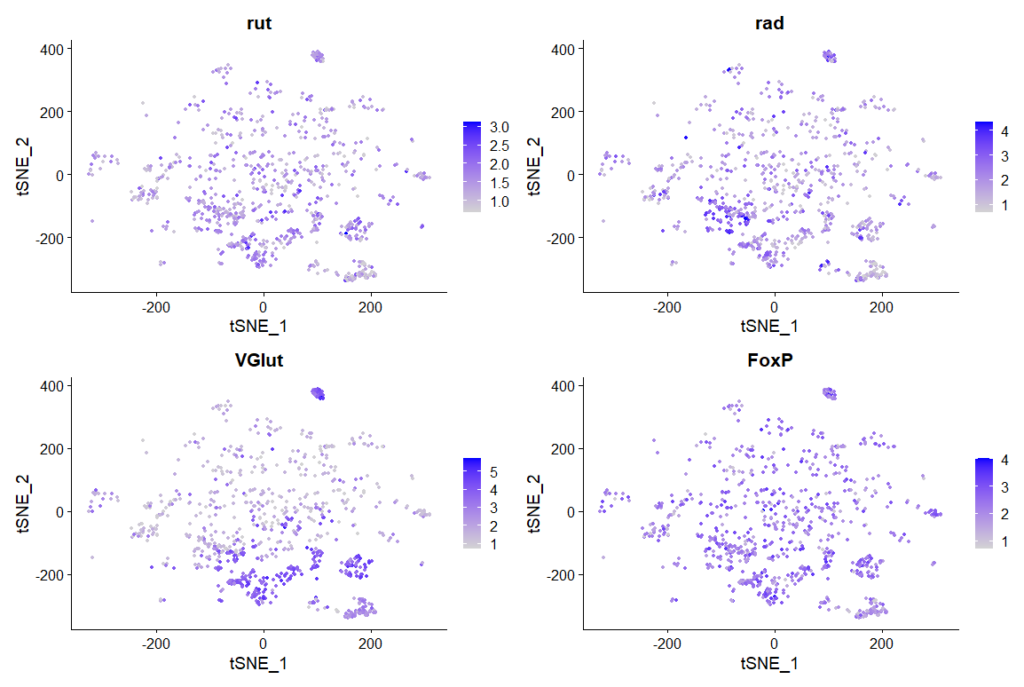
Top genes expressed in all of these cells:
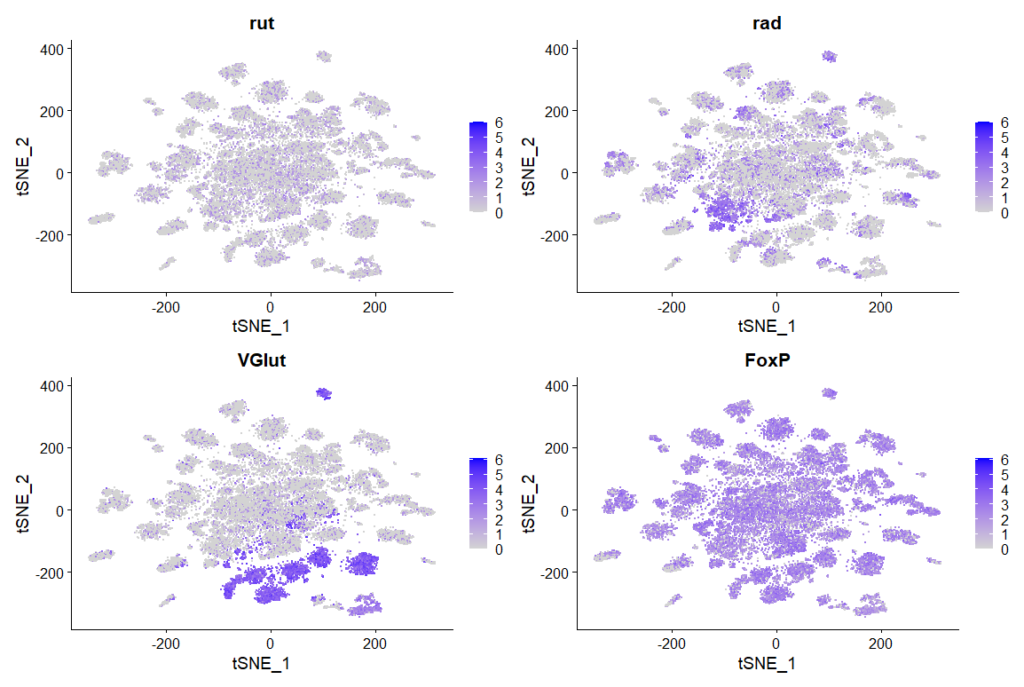
Rut and Rad Expression in Ventral Nerve Cord
I replicated the analysis of the single-cell RNA-sequencing data of the ventral nerve cord from Allen et al., (2020)
tSNE cluster map
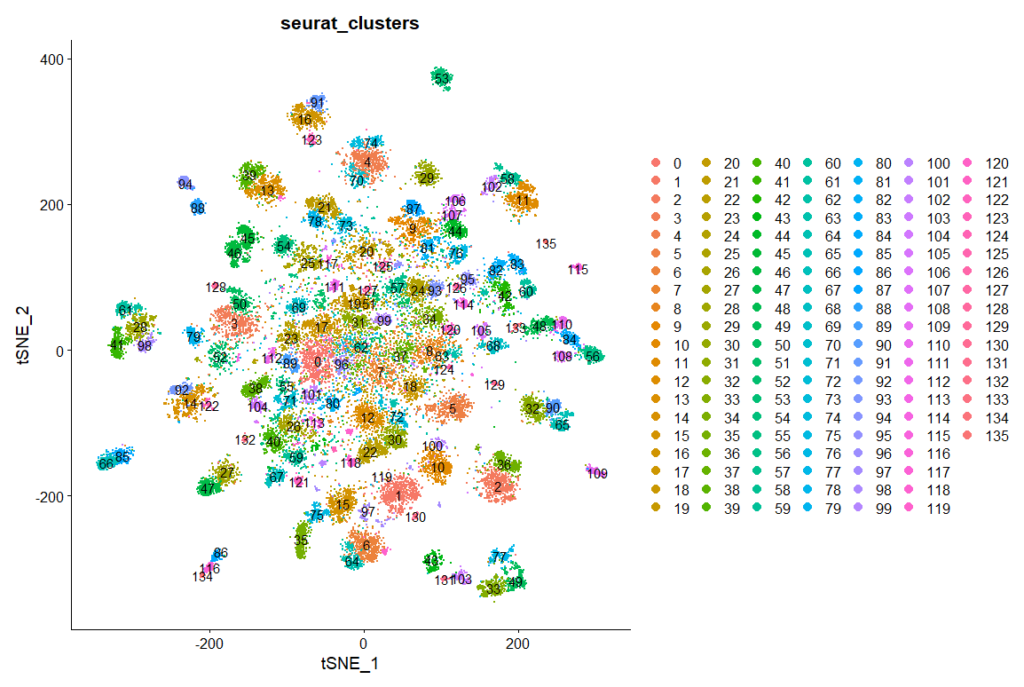
rut, rad, VGlut, FoxP expression in clusters:
Aaron M Allen Megan C Neville Sebastian Birtles Vincent Croset Christoph Daniel Treiber Scott Waddell Stephen F Goodwin
(2020) A single-cell transcriptomic atlas of the adult Drosophila ventral nerve cord
eLife 9:e54074.
https://doi.org/10.7554/eLife.54074Torquemeter Test and Confocal images of MBON02
16 flies went through the test out of 48 flies
I retook the images of last week’s brains for MBON02-transtango (trans-tango mkII) imaging. Green channel shows MBON02, red channel shows post-synaptic cells of MBON02:
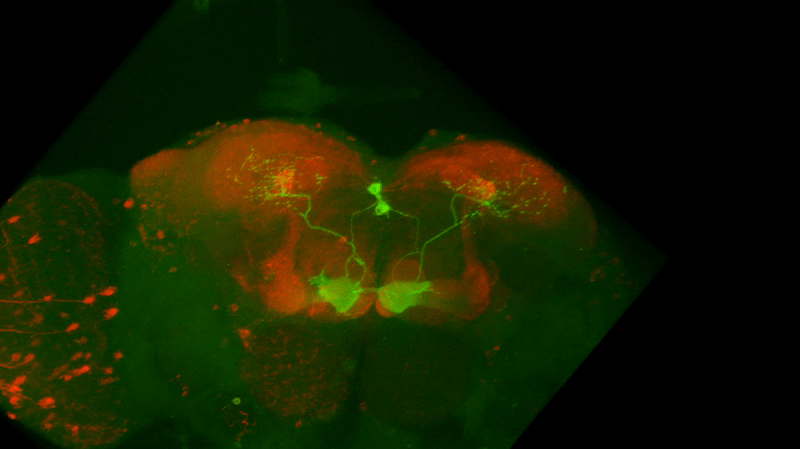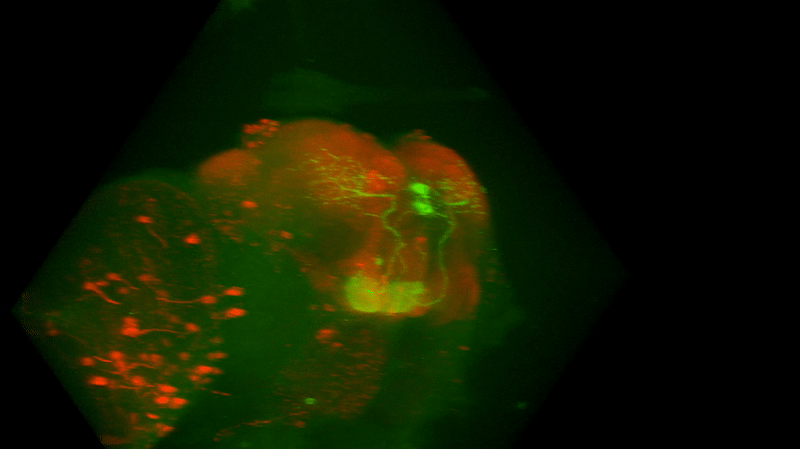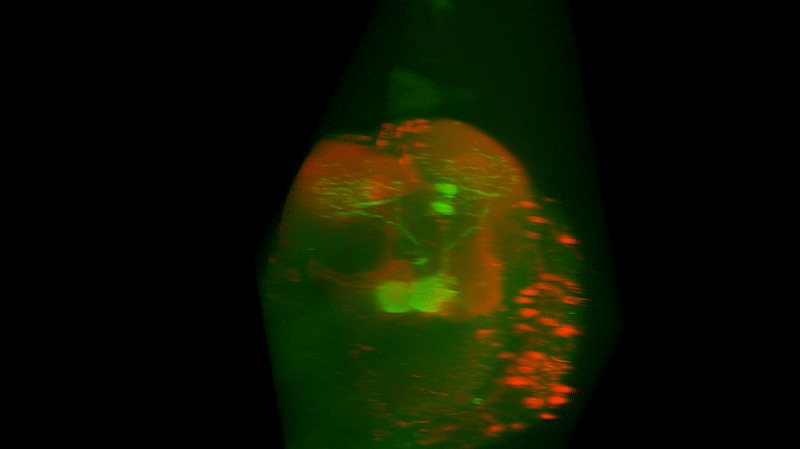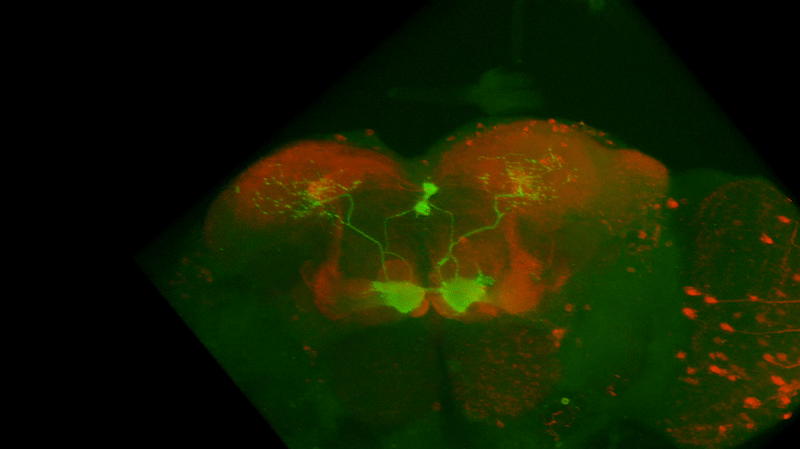
MBON02 Trans-tango images
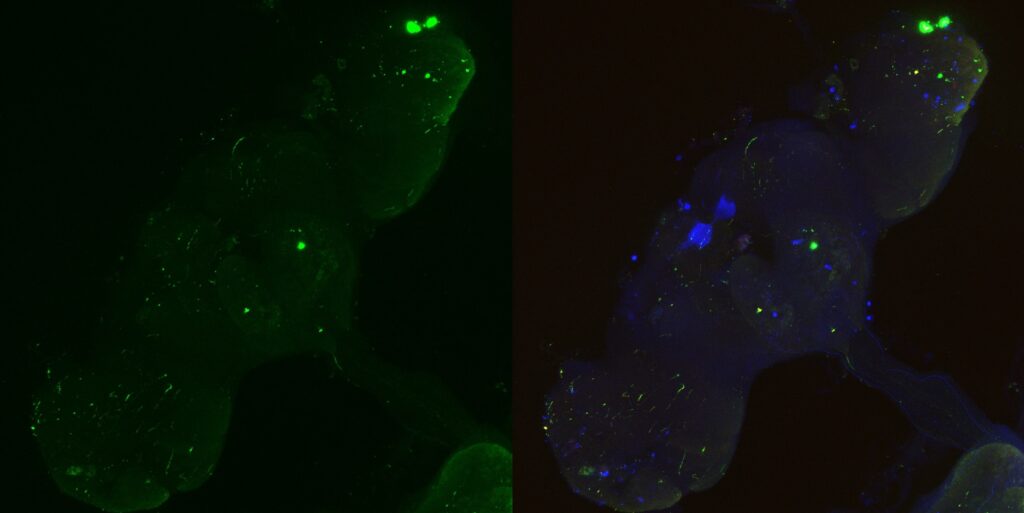
MBON02 trans-tango image from Kaun Lab. Left image shows trans-tango signal in red, right image shows overlay of all channels (green for MBON02, red for trans-tango, blue for nc82):
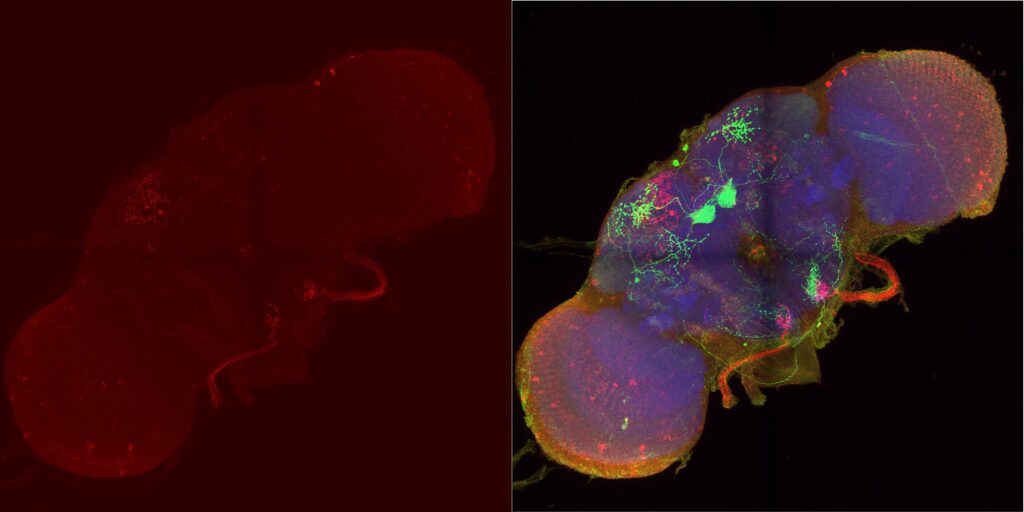
My MBON02 trans-tango images stained with anti-HA instead of anti-RFP. Flies generated by crossing trans-tango mkII line (BDSC: 95317) with MB399B (Brembslab stock number: 319). Left shows trans-tango signal in green, right shows overlay of al channels (green for trans-tango, blue for MBON02, red for nc82):
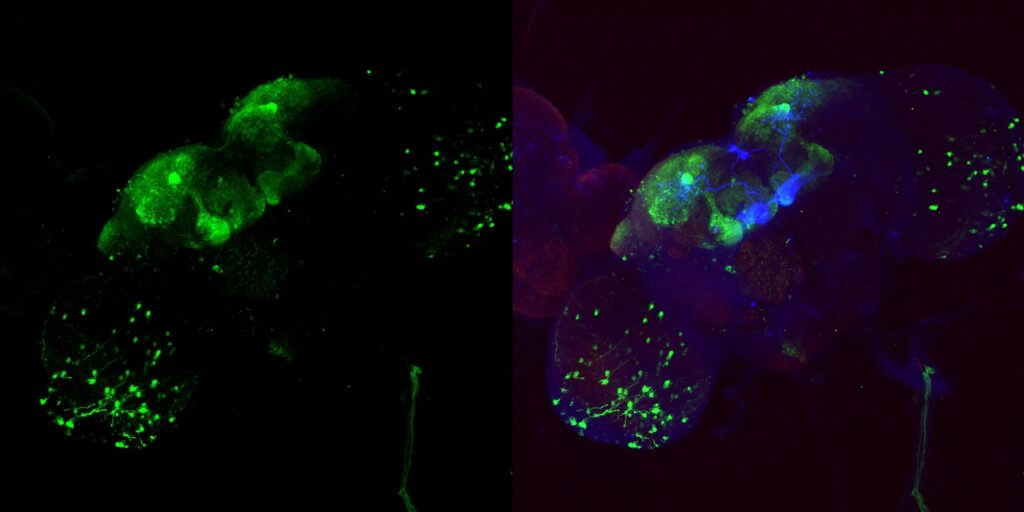
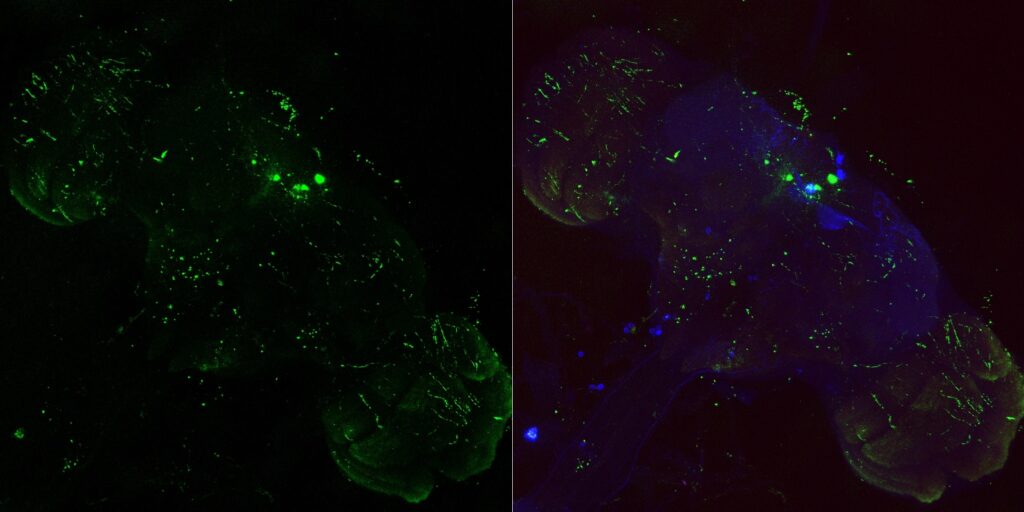
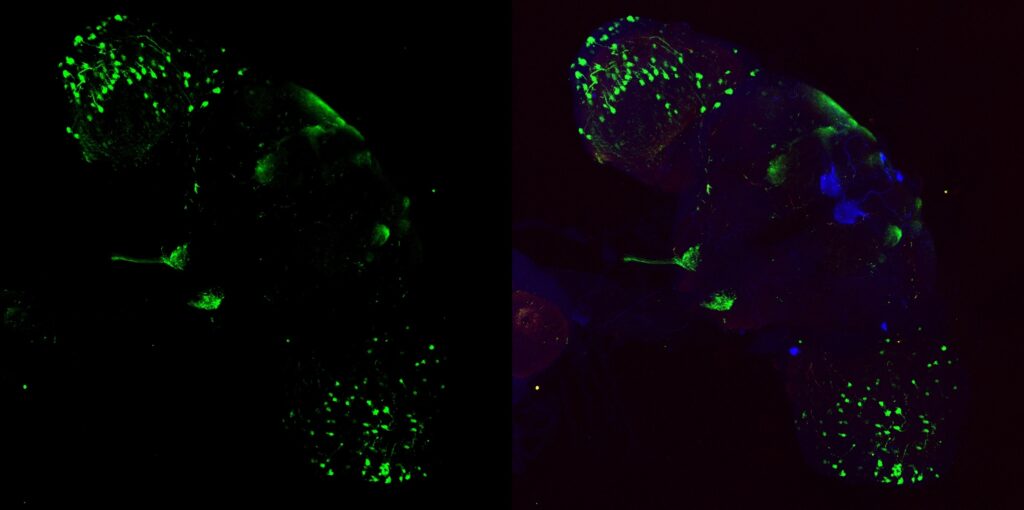
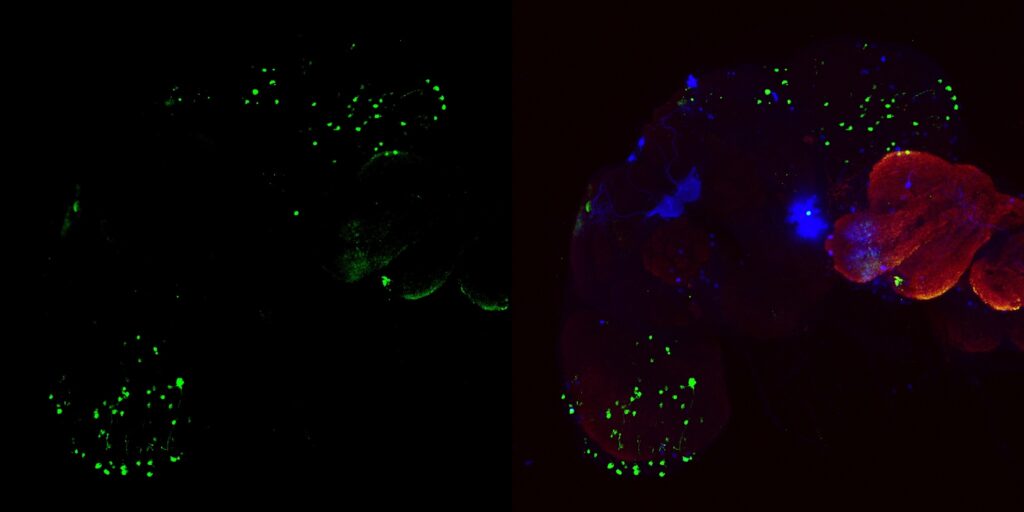
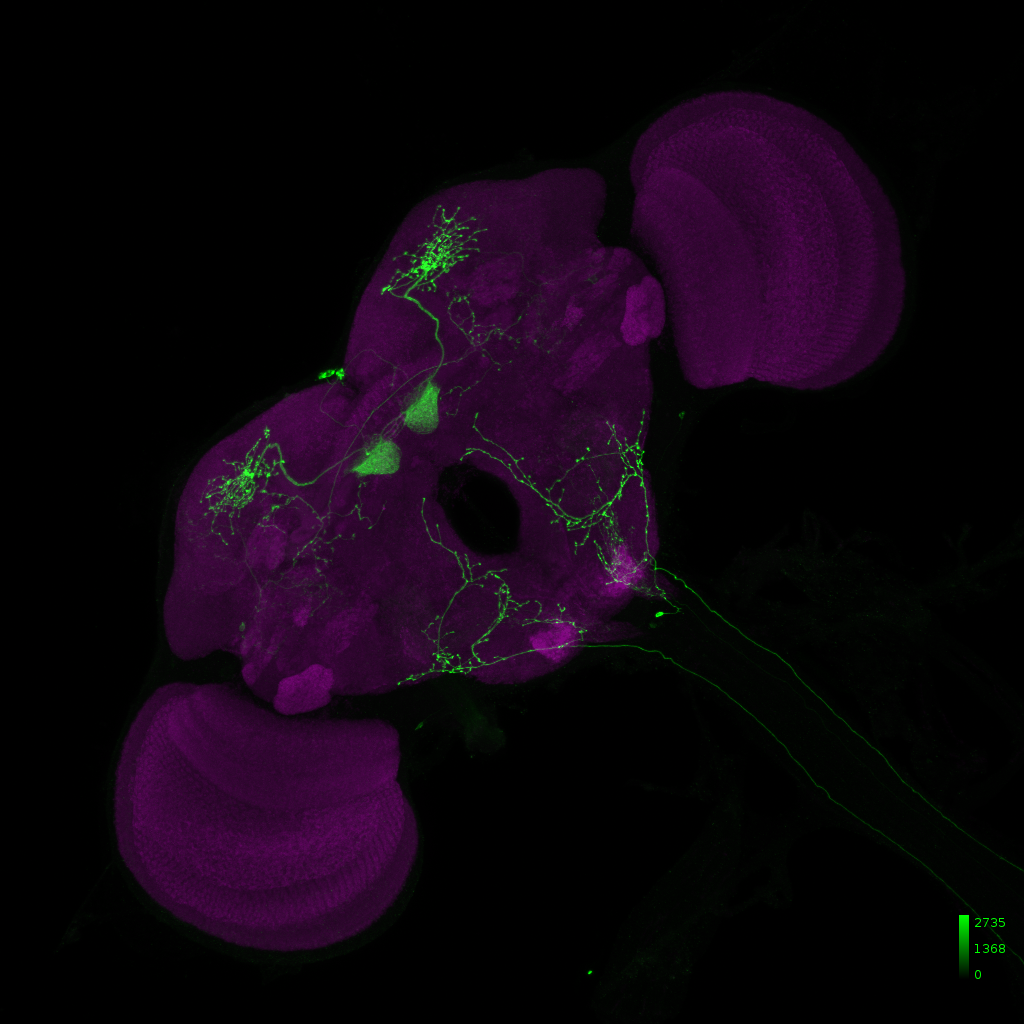
Janelia Flylight image of MB399B with 20XUAS-CsChrimson-mVenus trafficked in attP18 reporter:
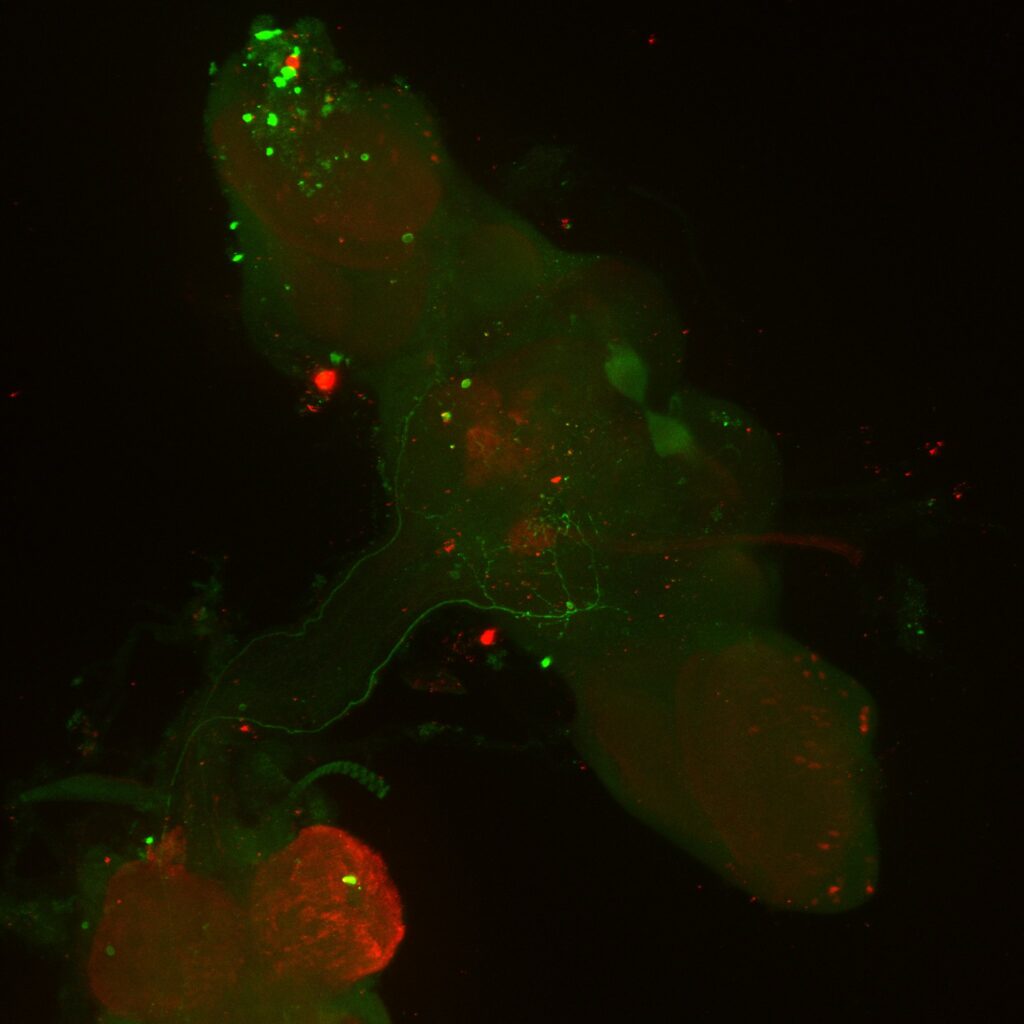
Confocal Images Trans-Tango, Retro-Tango
Staining done by anti-GFP (MBON02), anti-RFP (post-synaptic cells), nc82 (for background stainingm not shown here). MBON02 cells are clearly visible on green channel but there is no clear cell signal on red channel (MBON02 in green channel, postsynaptic cells in red channel).
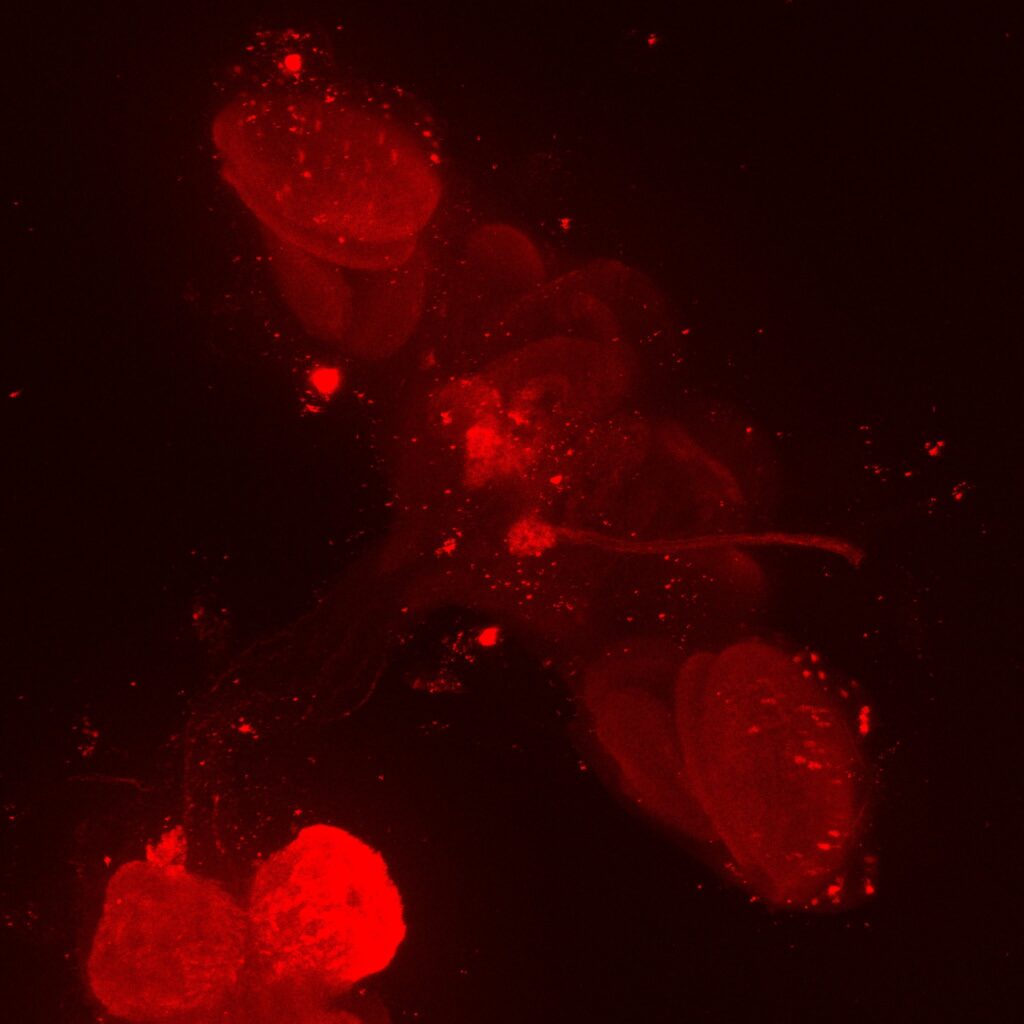
Here is the trans-tango image of MBON02 (postsynaptic cells of MBON02) from Kaun Lab:
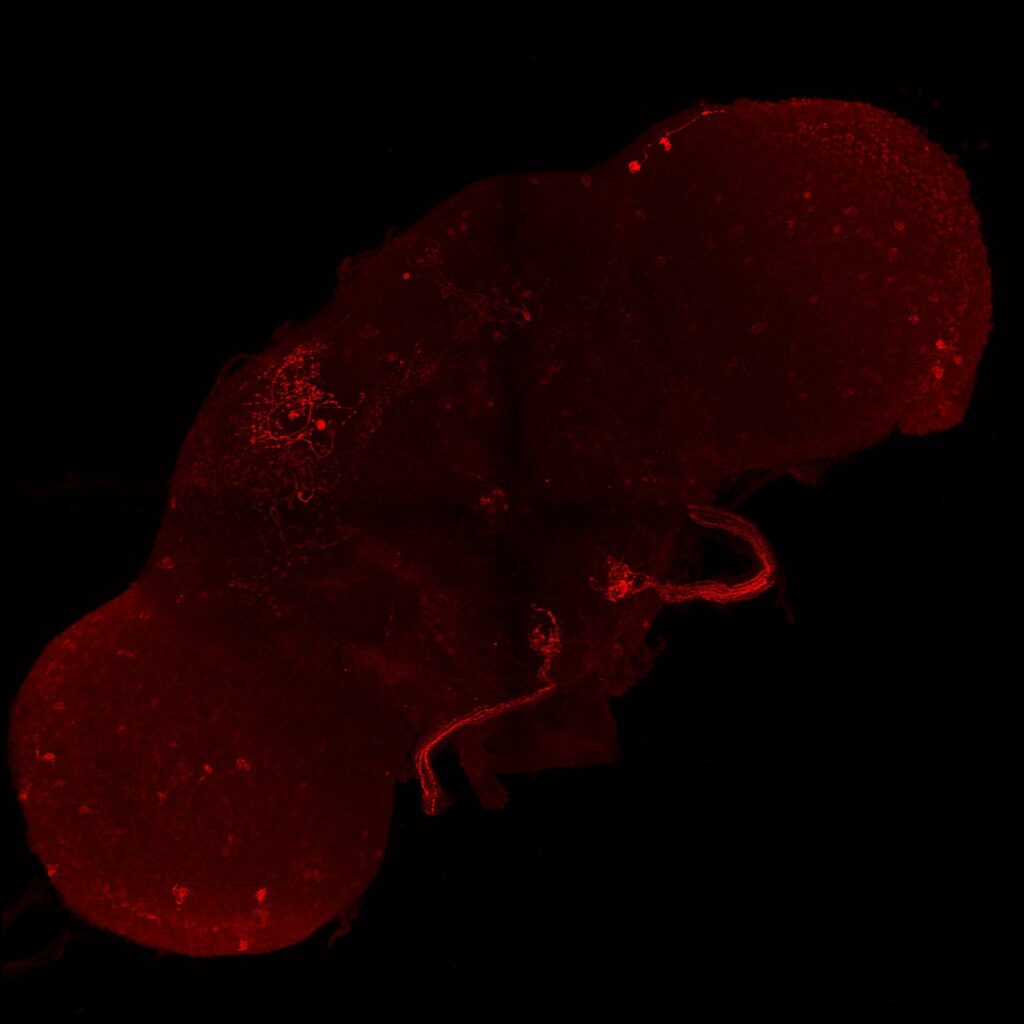
Retro-Tango of b3 mns look worse, nearly no cell signal both in green (b3 mn) and red channels (b3 mn presynaptic neurons)