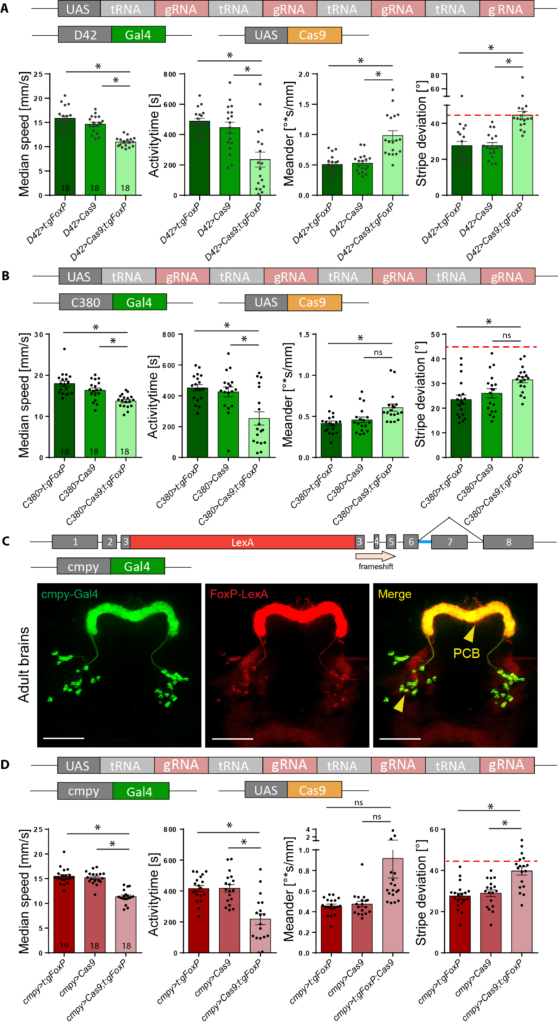
pCDF6 cloning with three inserts
on Sunday, November 8th, 2020 7:33 | by Sarah-Lynn Stratil
pCDF6 digestion with enzyme Bbs1
-5.5µl vector (undigested), conc.: 539.1 ng/µl
-1 µl enzyme (Bbs1)
-5 µl buffer (CutSmart)
-38.5 µl H2O
Agarose-gel electrophoresis
E.Z.N.A. Gel Extraction-DNA Purification from Agarose gel
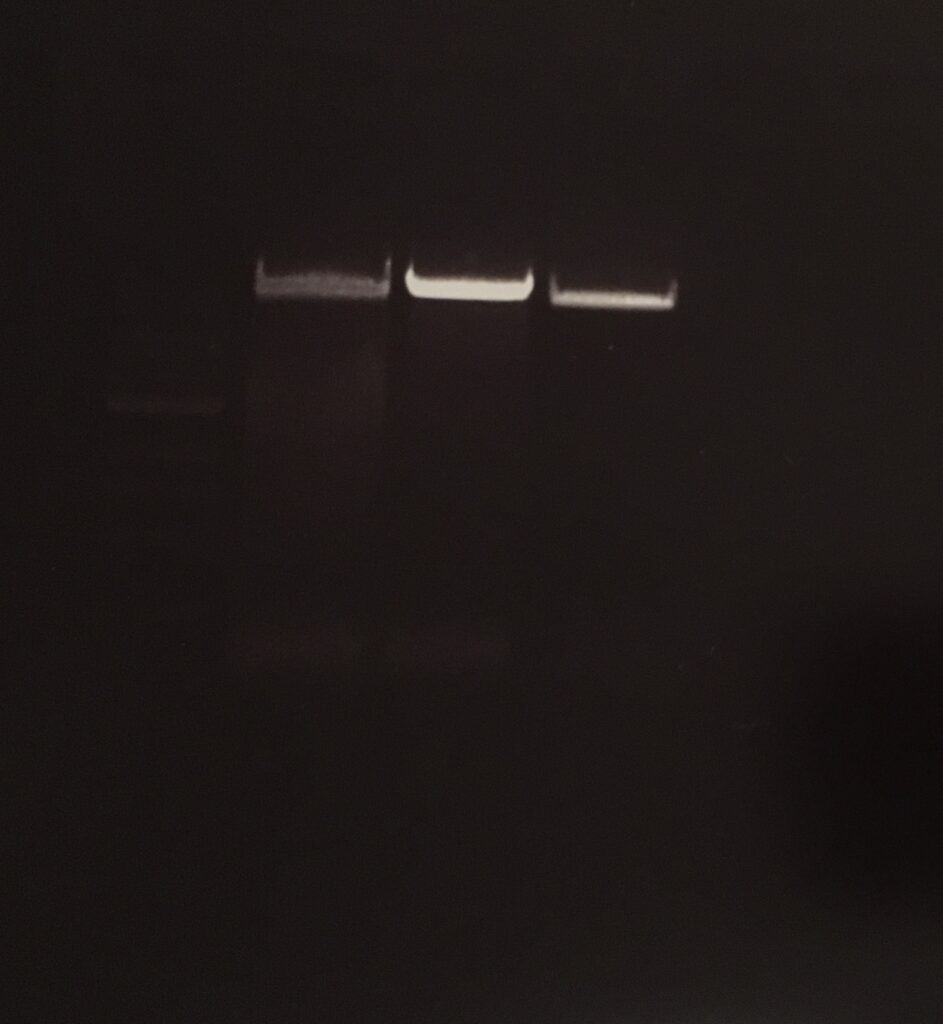
concentration: 38,0 ng/µl
PCR pCDF6 Primer
-12.5 µl primer reverse/forward
-5 µl dNTPs
-2.5 µl Taq-Polymerase
-0.29 µl pCDF6 digested
-50 µl buffer
-167.5 µl H2O
Agarose-gel electrophoresis
E.Z.N.A. Gel Extraction
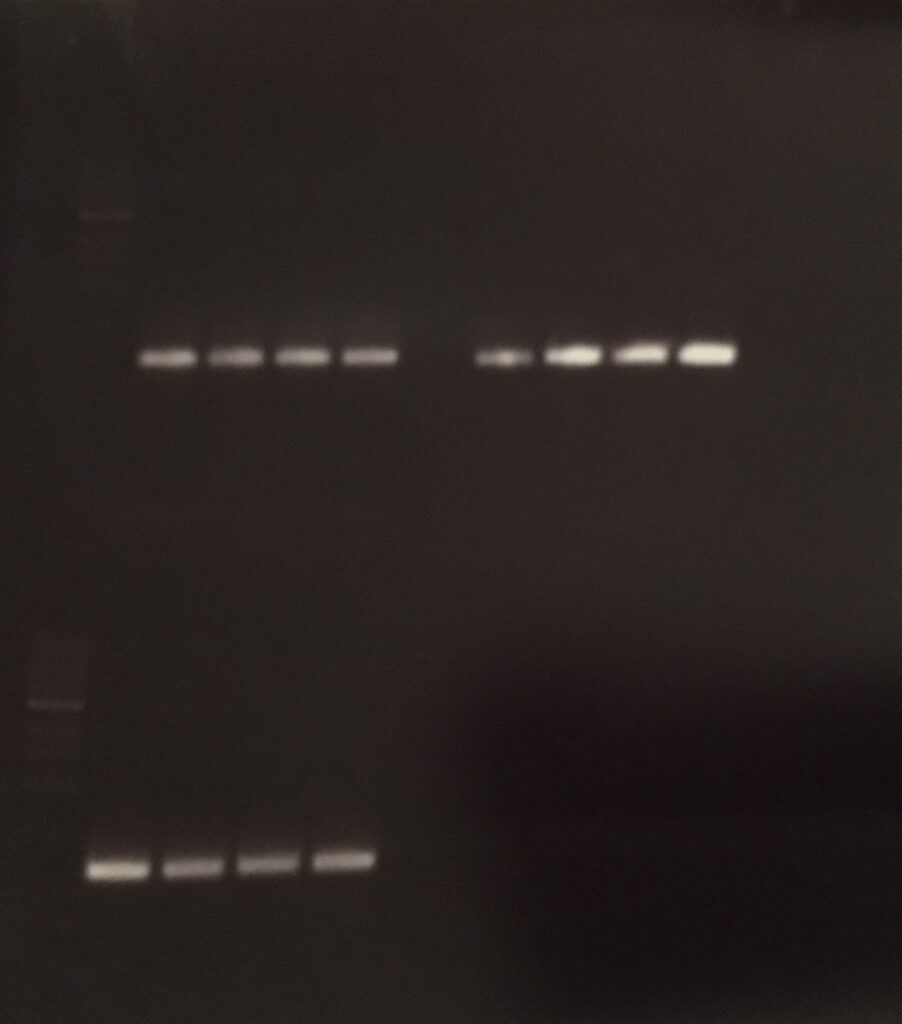
primer 1: 170.7 ng/µl
primer 2: 137.4 ng/µl
primer 3: 215.0 ng/µl
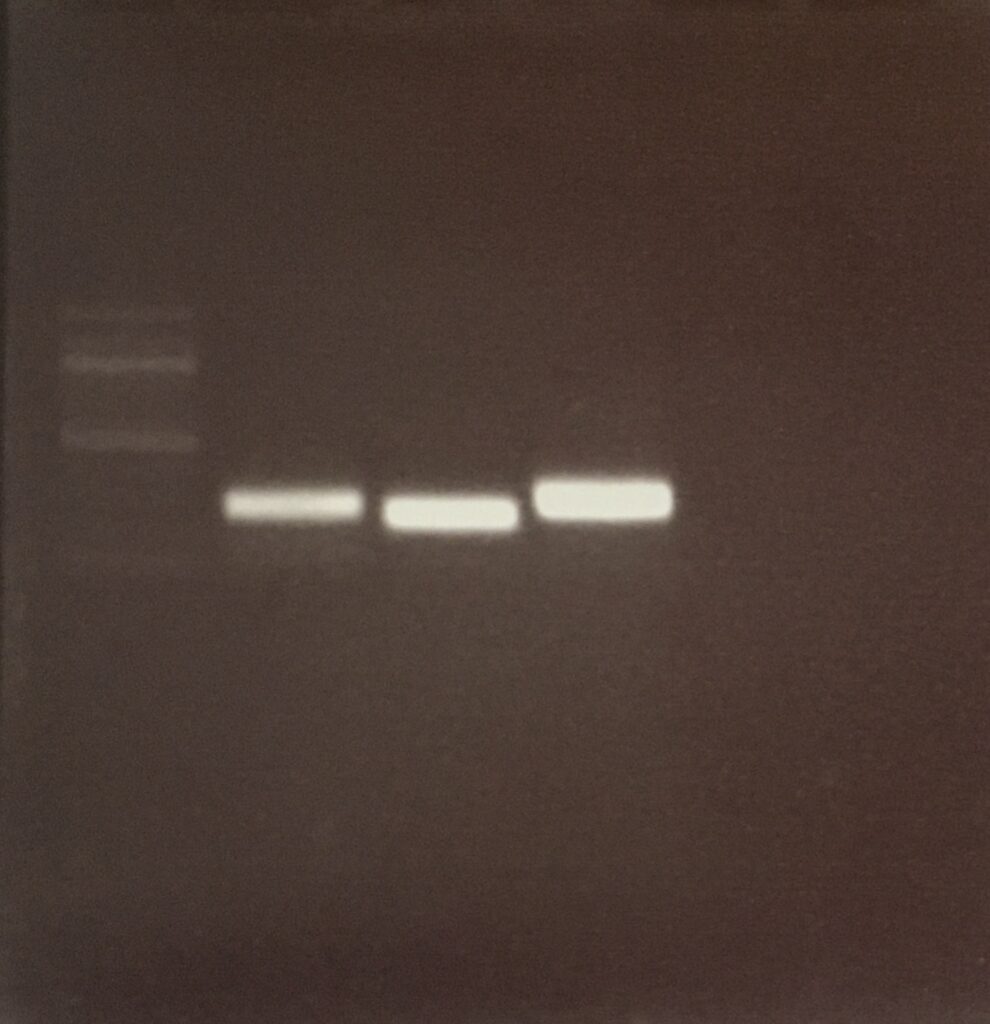
pCDF6 NEBuilder Assembly Reaction
-pcr1: 4.85 ng -> 1.60 µl
-pcr2: 4.24 ng -> 1.20 µl
-pcr3: 4.87 ng -> 1.20 µl
-pCDF6: 100 ng -> 2.90 µl
-H2O: 3.1 µl
-10 µl NEBuilder HiFi DNA Assembly Master Mix/Control
heat shock transformation
-> plated on LB0+Amp plates
-> no colonies
colony PCR (29.10.20)
-37 µl primer forward/reverse
-37 µl dNTPs
-37 µl Taq Polymerase
-74 µl buffer LSB
-518 µl H2O
Agarose-gel electrophoresis
colony pcr
Testing of possible Gal4-lines
on Monday, September 28th, 2020 12:11 | by Andreas Ehweiner
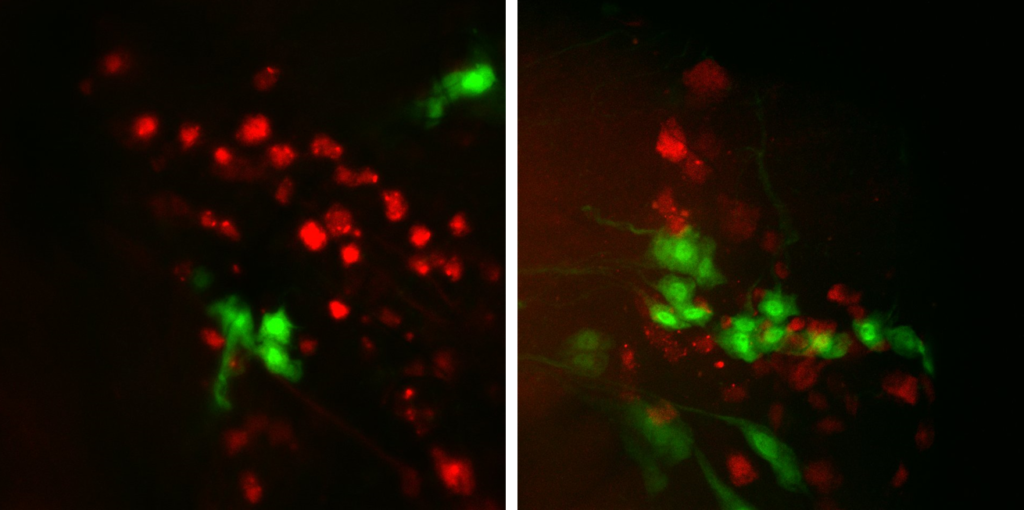
I finished my cross for the colocalisation of FoxP with the 6 Gal4-lines i ordered.
Two lines show a nice colocalisation and will be tested.

One would show some overlap, but a completly diffrent expression pattern than it should have. So i will try this line again from the stock to exclude a mixup of the line.
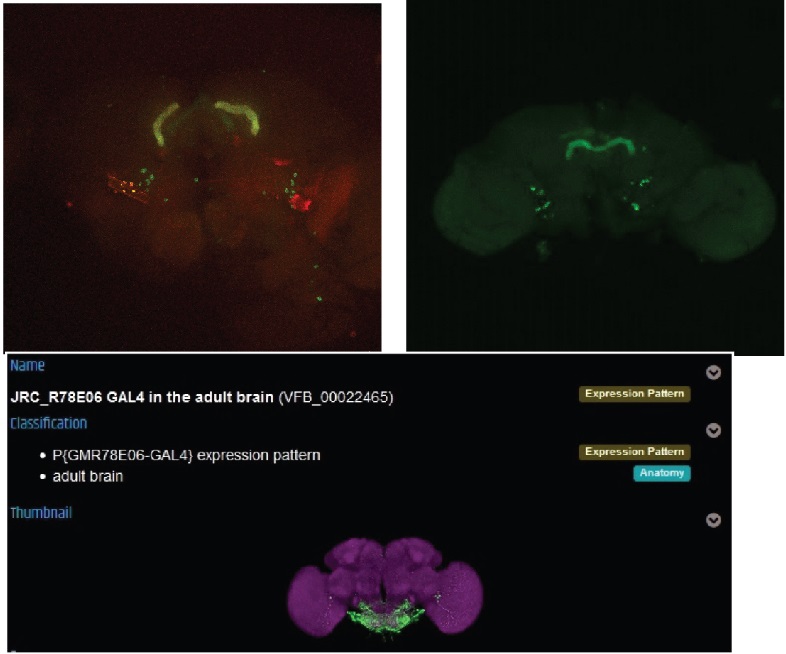
The next one dont seem to have any overlap, but also the pattern looks not like it should.
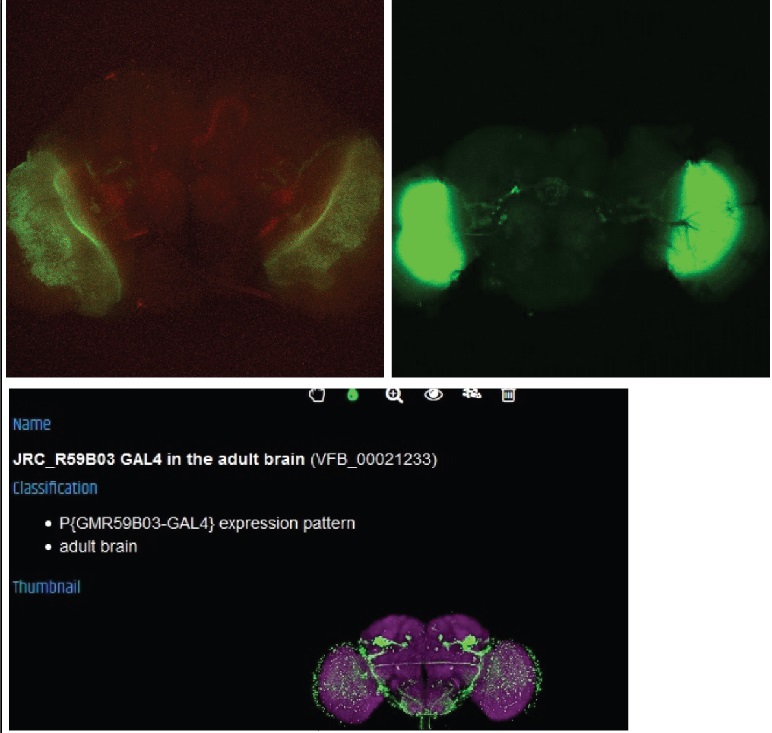
This line seems to have coexpression but the pattern also looks a bit odd, this may be due to a general weak signal and a bad dissection.
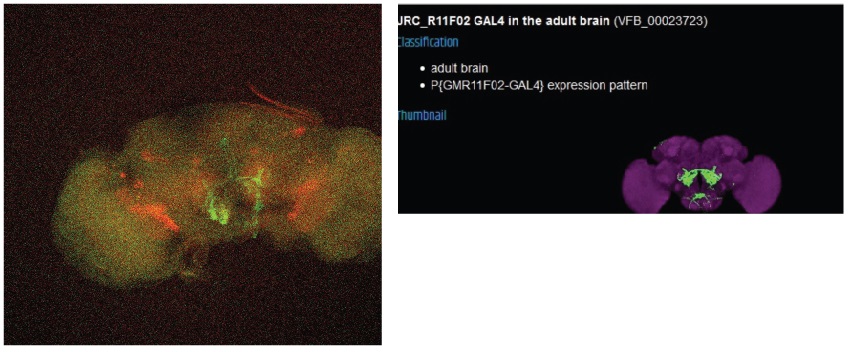
The final line seems to have no overlap with the FoxP expression.
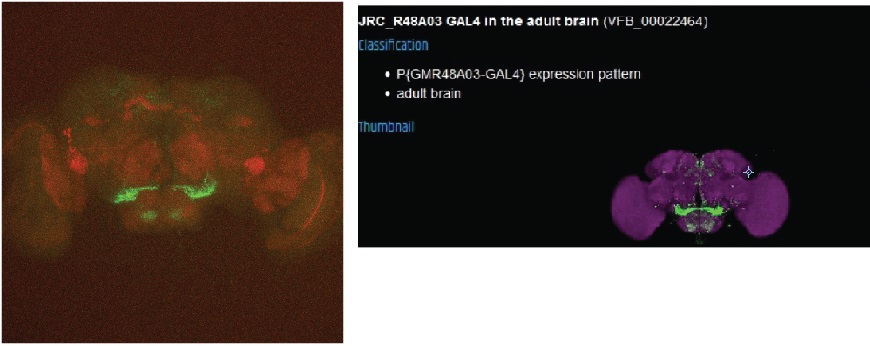
Category: Anatomy, crosses, Foxp | No Comments
FoxP knock out in adult.
on Monday, September 14th, 2020 1:49 | by Andreas Ehweiner
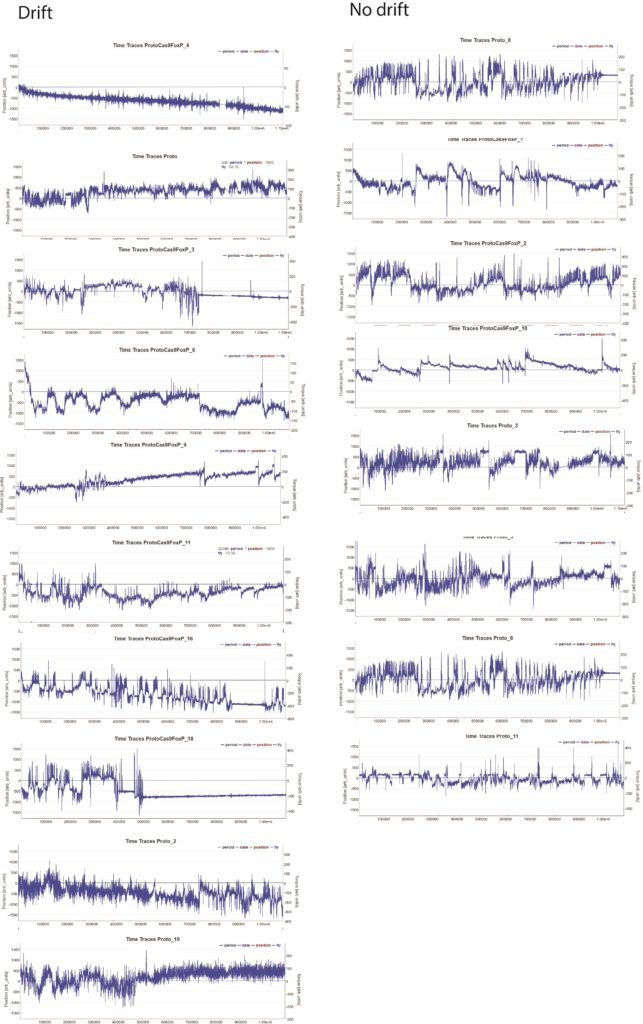
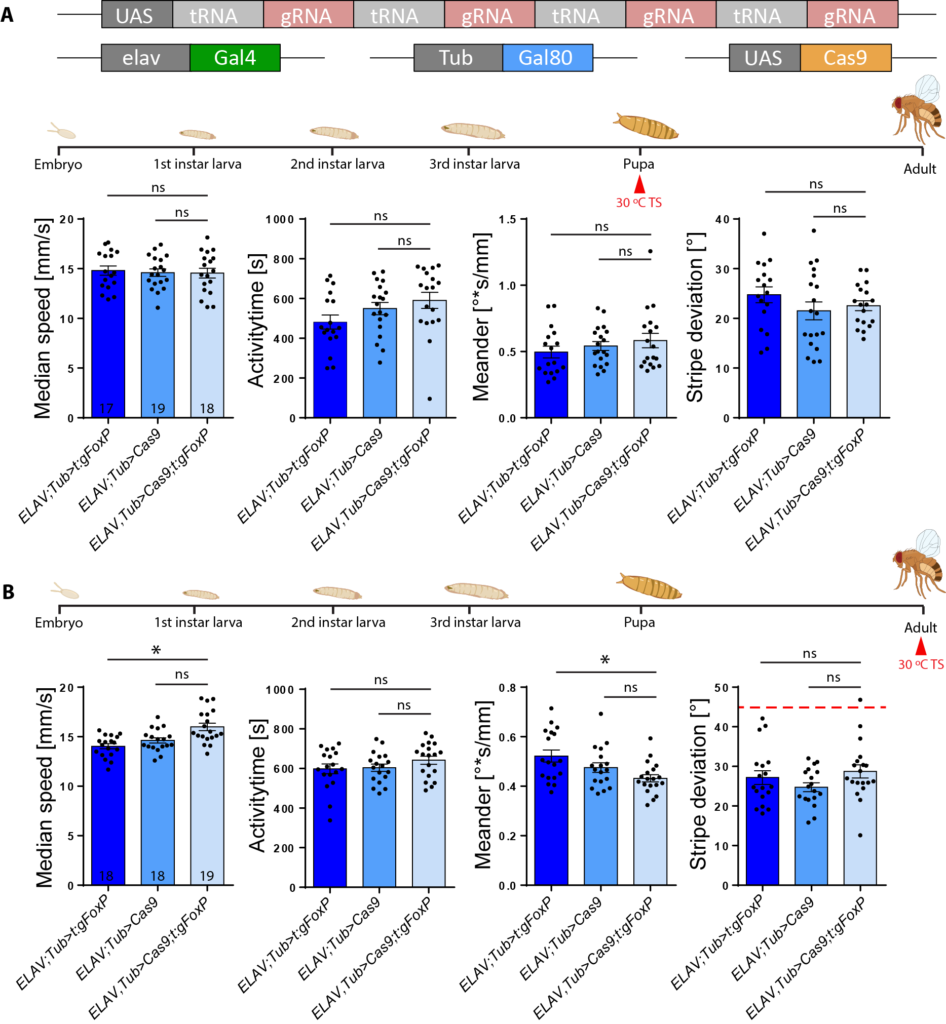
The repetition of the knockout experiment in adult flies indicates that FoxP is not required for learning in adult flies. (Exp = elav tubGal80>Cas9gFoxP)
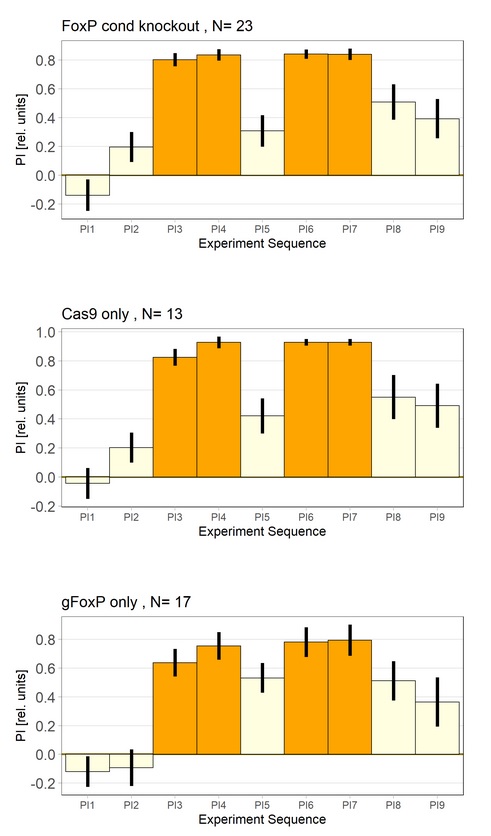
Category: flight, Foxp, Memory, operant self-learning | No Comments
PCB x Cas9gFoxP
on Monday, August 31st, 2020 8:47 | by Andreas Ehweiner
Knockout of FoxP in the PCB. Flies are still able to learn.
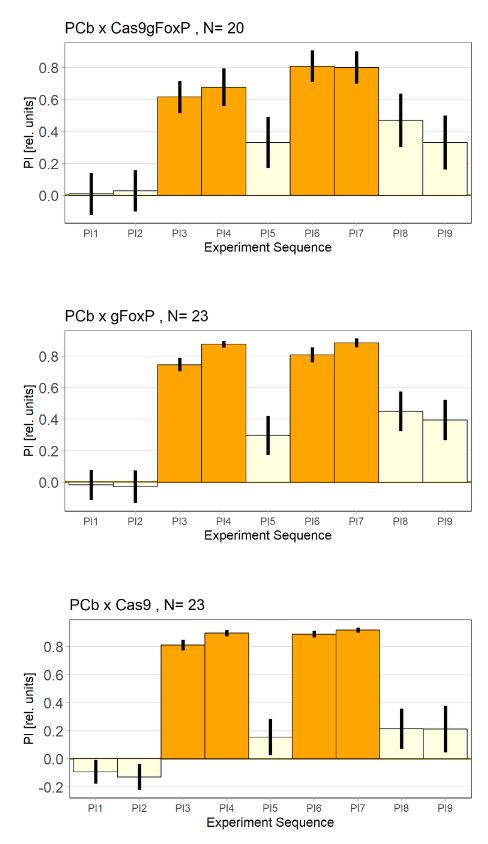
Category: flight, Foxp, operant self-learning | No Comments
August
on Thursday, August 27th, 2020 1:46 | by Ottavia Palazzo
- Halfway in analyzing the Buridan data altogether (not in two batches)
- Maxiprepped plasmid for making FoxP protein at the klinikum
- Writing the thesis a little bit
- bought Fas-II antibody
________________________________________________________
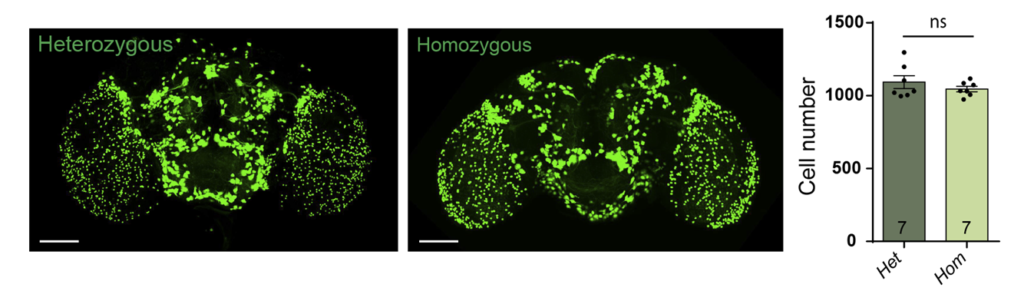
- data 1): FoxP-iB Heterozygous/Homozygous comparison with Stinger-GFP. This time i was cautious with everything: fly all the same age and sex, same larvae density, same number of copies of GFP
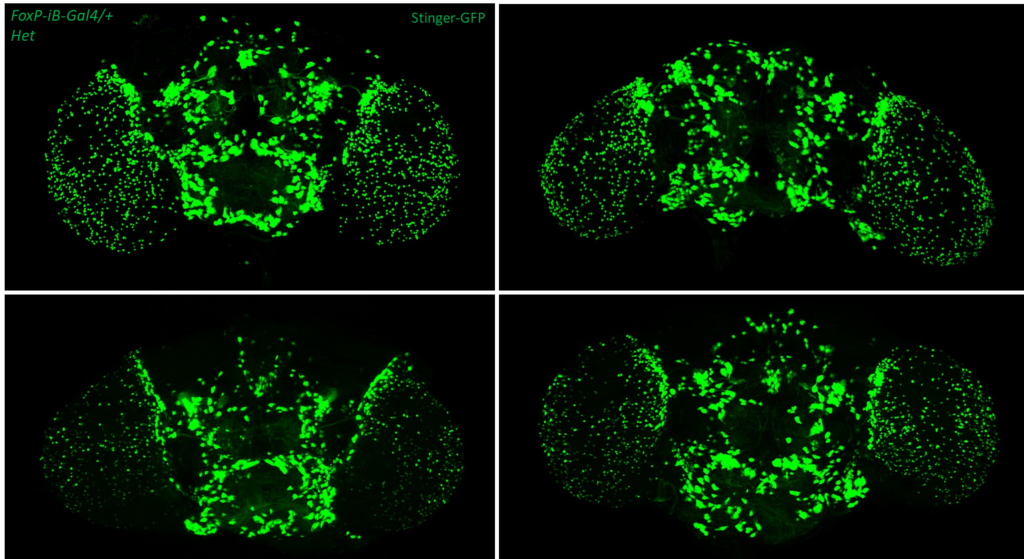
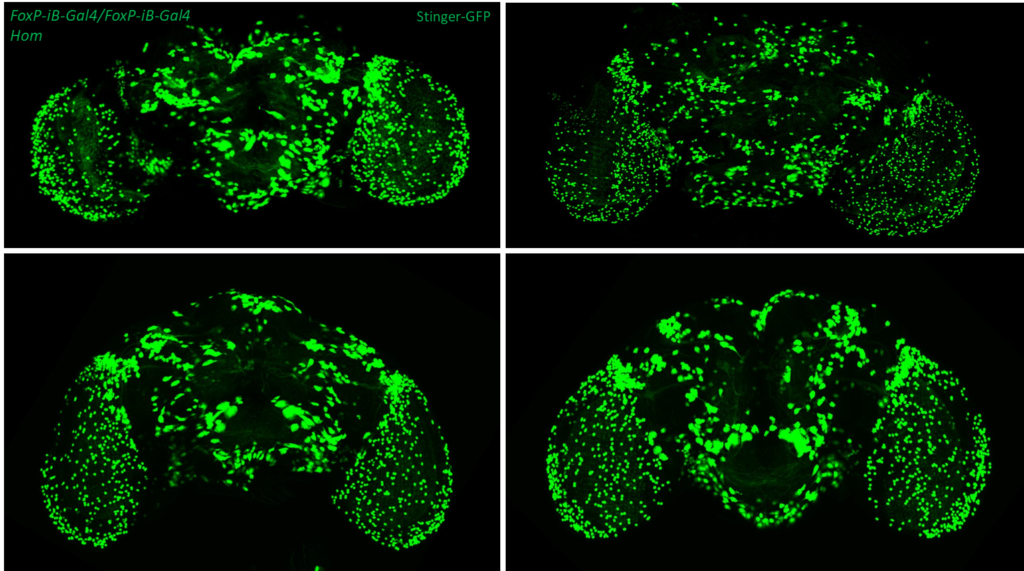
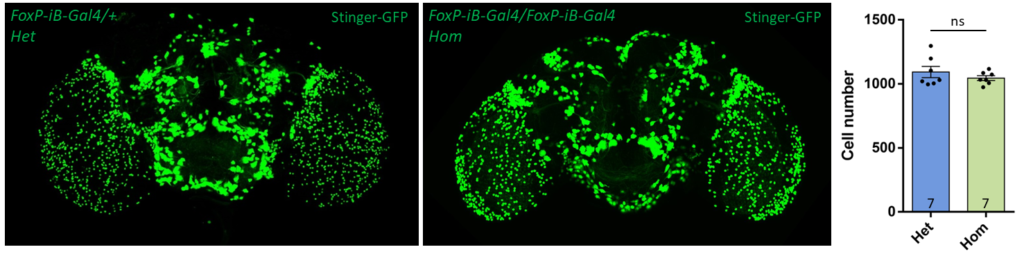
I can not detect any difference between homozygous and heterozygous mutants. I also counted cells in IMARIS and I detected no difference in number or distribution.
- data 2): FoxP-iB Heterozygous/Homozygous comparison with CD8-GFP. This time i was cautious with everything: fly all the same age and sex, same larvae density, same number of copies of GFP
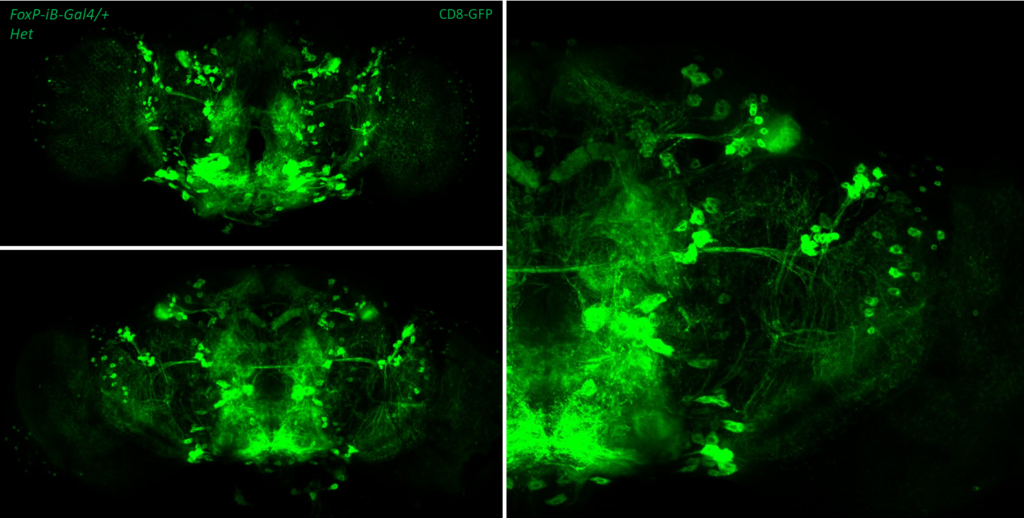
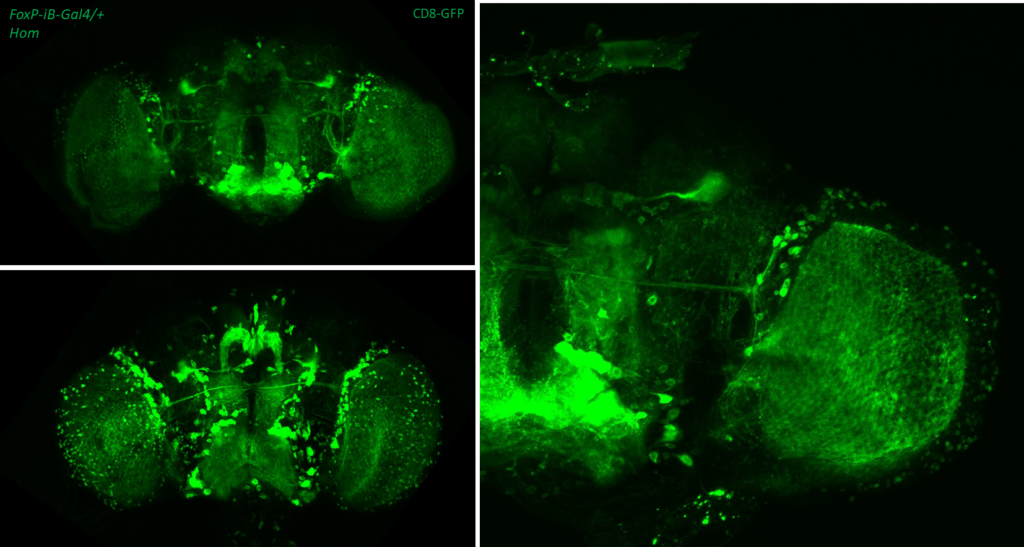
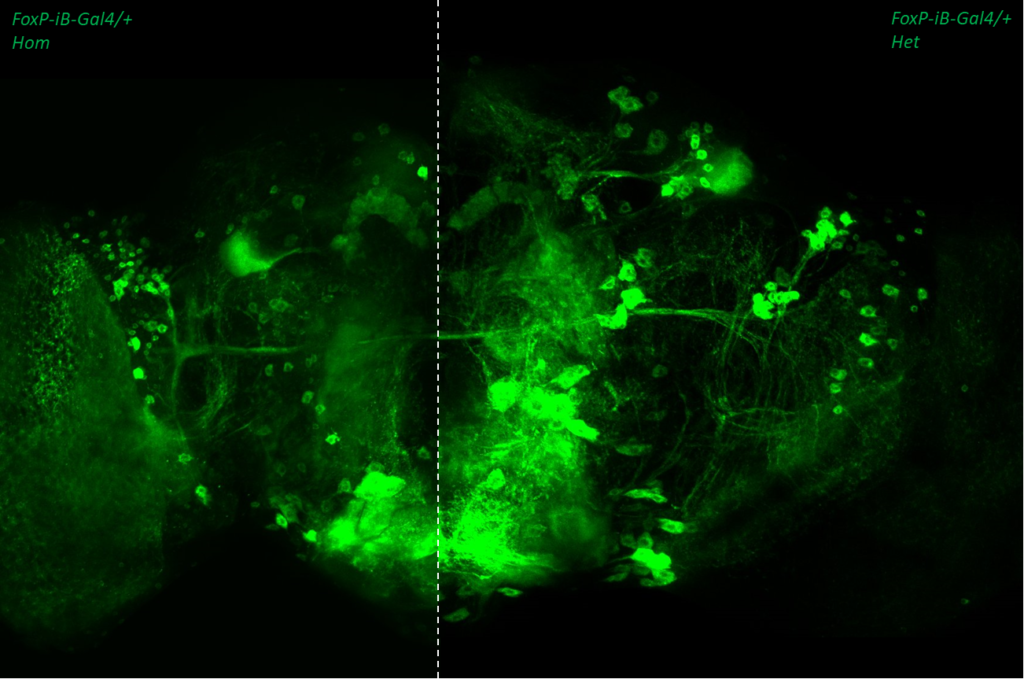
I can see some differences but i do not know how to quantify/explain them. Also I am not sure if it is a problem of dissection/mounting. I am not convinced.
- data 3): nc82 staining on homo/hetero FoxP-iB. I do not see differences

Problem: I haven’t managed yet to make the FasII antibody work:
I have used a 1:10 concentration of primary antibody for one night and a 1:100 concentration of secondary antibody for 4 hours. I will ask someone
In the picture I have increased the gain a lot just to be sure. I could not see anything
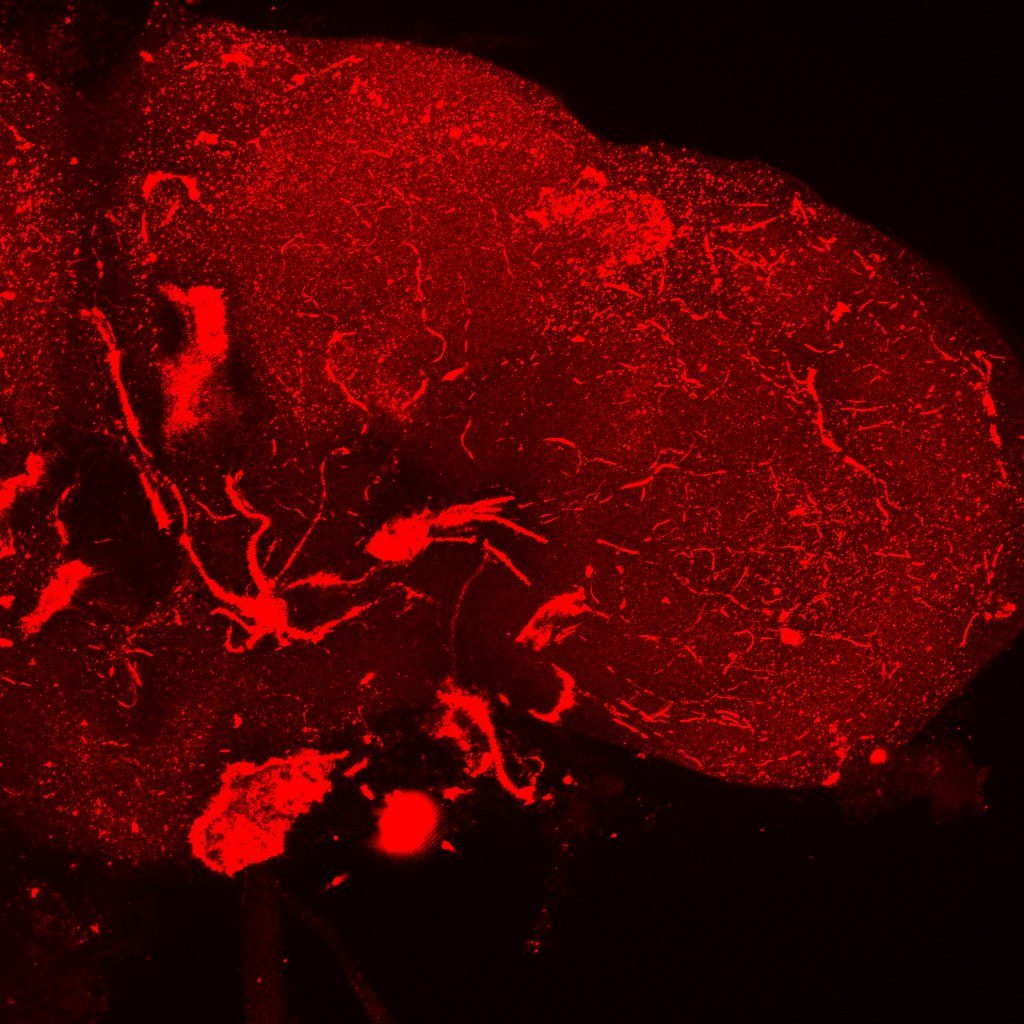
Category: Foxp | No Comments
Recent experiments
on Monday, June 22nd, 2020 1:52 | by Andreas Ehweiner
Category: flight, Foxp, operant self-learning, set-up test, Uncategorized | No Comments
Pupae TS increased N
on Monday, June 22nd, 2020 10:34 | by Ottavia Palazzo
Category: buridan, Foxp | No Comments
Tdc2-Gal4 + FoxP-LexA in VNC
on Thursday, June 18th, 2020 4:16 | by Ottavia Palazzo
Category: Foxp | No Comments
C380-increased N
on Wednesday, June 17th, 2020 9:49 | by Ottavia Palazzo
Category: buridan, Foxp | No Comments