Better analysis of slope dependence on flight length and further insights in the SMAP code
on Monday, December 4th, 2017 1:02 | by Christian Rohrsen
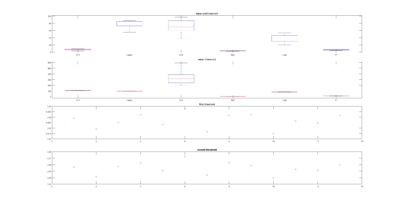
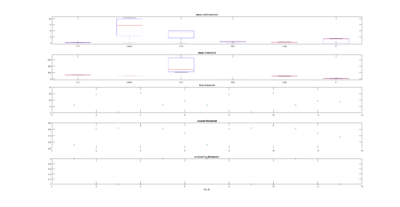
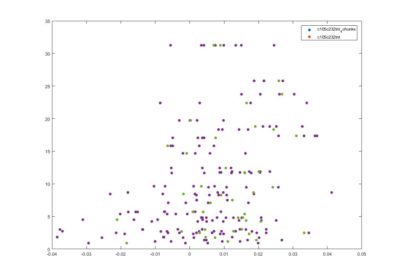
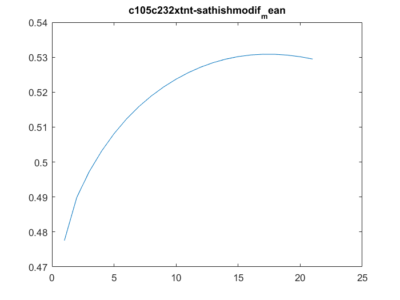
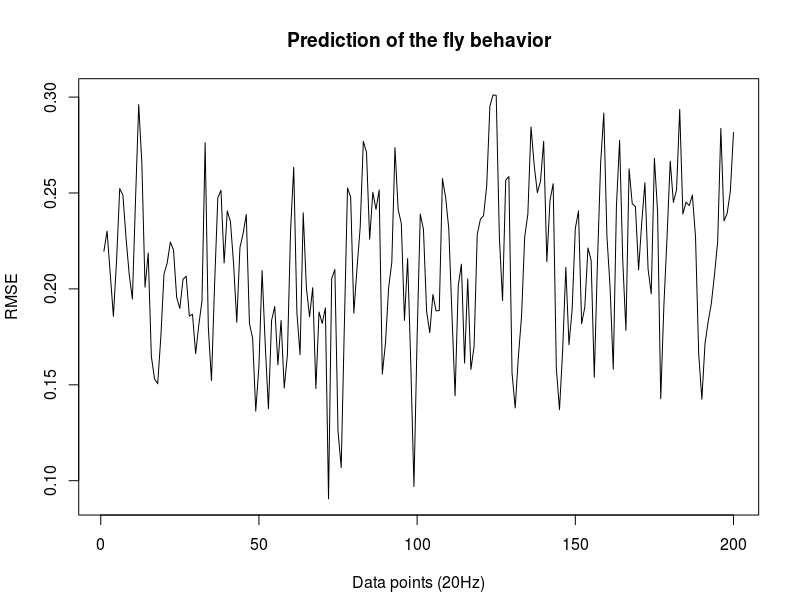
This is another way of showing how the slope of the SMAP analysis varies with length. I have choped the time series in 4 chunks and saw what was the slope for the chunks and for the whole time series. I think this is the best statistical way of doing it, it should not depend on what line was tested. I cannot see any effect.
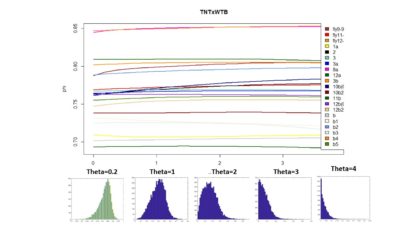
Here below is just to show what I found out in the code. What I thought that theta was controlling for nonlinearity in the model, for me it seems rather a control for under- overfitting in the model. So I might need to read the paper and see what do they say, and if it is the same as what I found out in the code.
Category: flight, Spontaneous Behavior, strokelitude, WingStroke | No Comments
Do not agree with Sathish?
on Monday, November 27th, 2017 2:43 | by Christian Rohrsen
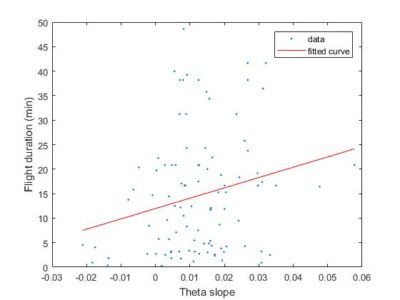
So this is the final graph assuming that 20Hz is the sampling rate. Sathish was not sure what it was and he said he will check.
To have a better overview how the length of the flight compares with the slope obtained from the SMAP. There is no big correlation whatsoever with around 100 flies. Sathish found this in his thesis with data from seventy something WTB, but to me it seems like an anecdotal result.
Here the same as above, just for showing the fit line.
Category: flight, Spontaneous Behavior, strokelitude, WingStroke | No Comments
All sathish data analysis
on Monday, November 20th, 2017 1:47 | by Christian Rohrsen
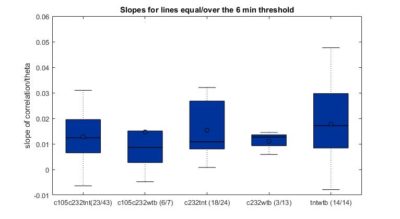
This are the results of all of the flies analyzed from Sathish. I just need to know the frequency of the acquisition to exclude the ones that are below 6 minutes.
Category: flight, Spontaneous Behavior, strokelitude, WingStroke | No Comments
Sathish data
on Monday, November 6th, 2017 1:37 | by Christian Rohrsen
I have analyzed the c105+c232 > tnt data from Sathish. There are a total of 43 flies, although many of them only flew for a few minutes, and therefore should be discarded. Below all of the individual fly scores. I need to analyse now the other groups. This is btw the modified data set, whatever that means for Sathish.
This is a video of the projection from the torque data from Maye et al. 2007. The spikes are not that well sorted in this case as in the Strokelitude. I guess this is because the spikes do not look so smooth.
Category: flight, Spontaneous Behavior, strokelitude, WingStroke | No Comments
Recurrence quantitative analysis
on Monday, October 30th, 2017 11:43 | by Christian Rohrsen
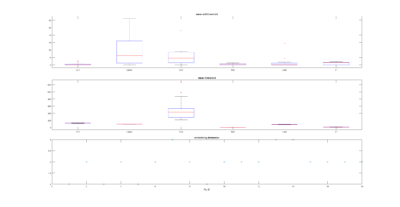
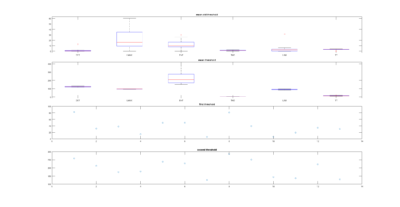
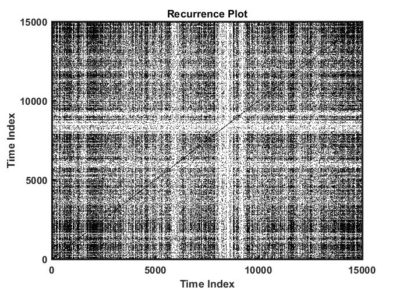
This is an example of a recurrence plot analysis. In the first graph is shown in single point in time in the optimal embedding dimension and the distance to the other points. For the recurrence plot analysis it is needed to put a threshold to make it binary. This is the second graph. From this second graph one can count many parameters like determinism, laminarity and so on. From what I see, the plots from the Strokelitude as well as Bjoern´s flight simulator in Maye et al 2007 show similar pattern (kind of crosses with vertical and horizontal lines).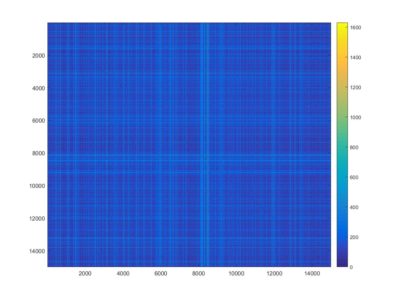
This is a measure of the Recurrence Quantitative Analysis of different groups. Recurrence threshold is a tricky and to some extent subjective measure, so this is why I tried two different ones.
DET: recurrence points that form a diagonal line of minimal length, the more diagonal, the more deterministic.
LMAX: Max diagonal line length or divergence. Sometimes considered as an estimator of max. Lyapunov exponent
ENT: Shannon entropy reflects the complexity of the system
TND: info about stationarity (trend)
LAM: Laminarity is related to laminar phases in the system (intermittency). It is tallied as vertical lines over a threshold.
TT: Trapping time, measuring the average length of vertical lines. Related to laminarity.
Automat
Openloop
Uniform
Category: flight, Spontaneous Behavior, strokelitude, Uncategorized, WingStroke | No Comments
More on the attractor
on Monday, October 16th, 2017 2:26 | by Christian Rohrsen
Video of the attractor projected from a chunk of flight trace. Here one can see the difference in the trajectories of up- and down spikes. So here is another way of spike sorting by the way :).
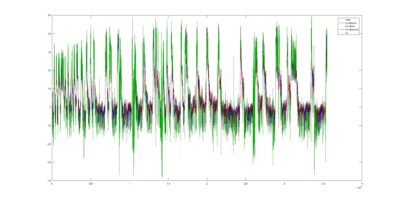
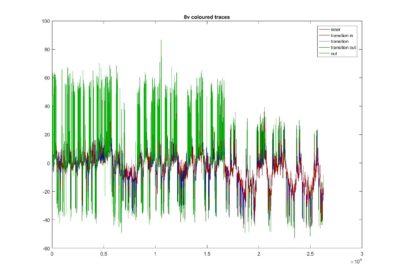
Coloured traces depending where the locate in the attractor projection. Green means that they lie outside and when they stay with the PCs around zero its in red. The fly1r above does not look so clean as the v8 fly below (the cleanest measure I have)
Category: Spontaneous Behavior, strokelitude, WingStroke | No Comments
SMAP results
on Monday, October 9th, 2017 2:34 | by Christian Rohrsen
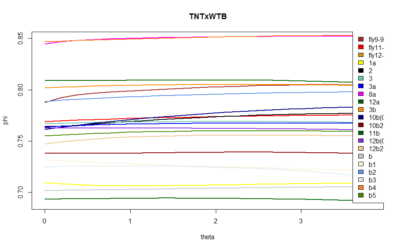
These are the results of the SMAP for the TNTxWTB. I also have done a few for the c105;;c232xWTB but there is not much to say. I would say that the cleanest lines show a bigger slope, but prone to subjectiveness.
In addition, I have done some animations of the attractors that I have posted on slack because of size.
Category: flight, genetics, R code, Spontaneous Behavior, strokelitude, WingStroke | No Comments
Attractors for c105;;c232>TNT
on Monday, October 2nd, 2017 5:35 | by Christian Rohrsen
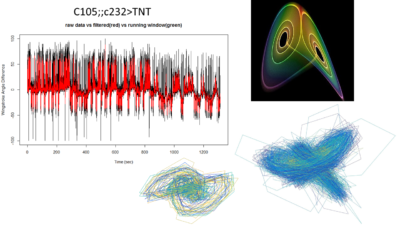
This is what I showed about one fly from this line showing the attractor.
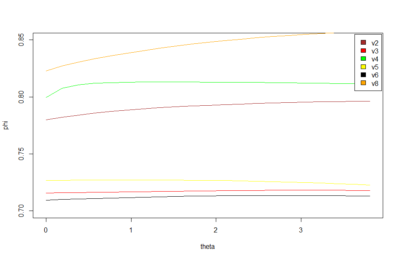
This graph is what I forgot to show in the lab meeting. There are the 6 best traces from this same line. All of them selected ad hoc subjectively. The three best of them to my eyes are exactly the three above in the graph (v8-this one is the one shown in the picture with the attractors-,v4,v2). What does this means? the ones with better traces (subjectively) have higher offset in the phi, this means that they are more predictable overall (maybe because more resolution?). In addition, they show higher slopes which means more nonlinearity.
Category: flight, Spontaneous Behavior, strokelitude, WingStroke | No Comments
Cumulative bins, starting at zero and normalizing
on Monday, April 4th, 2016 3:04 | by Christian Rohrsen
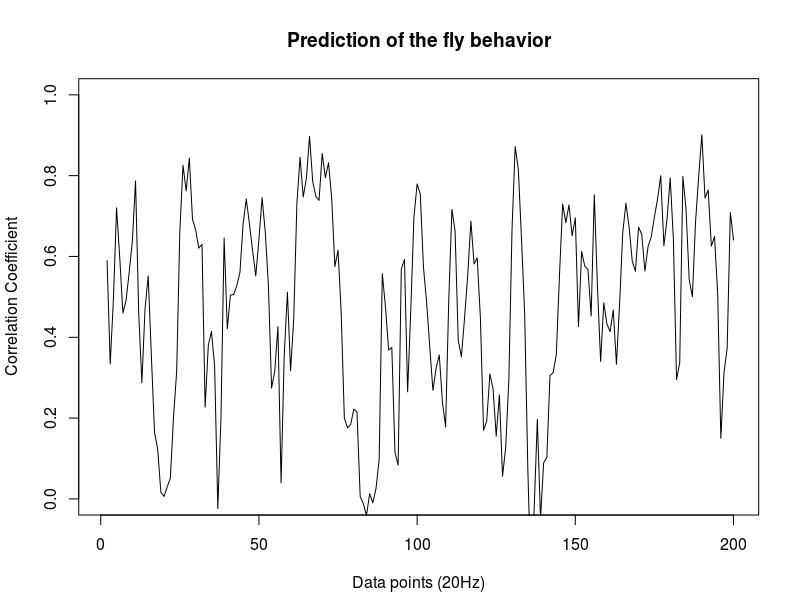
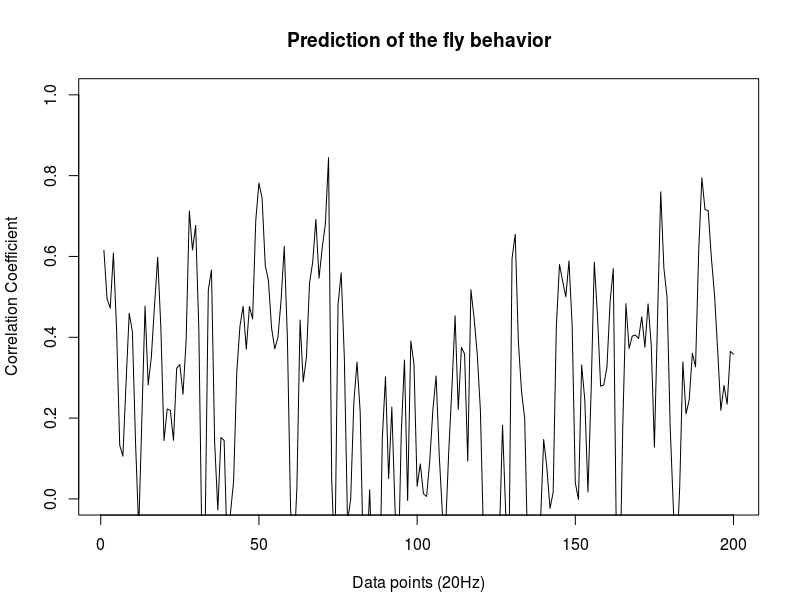
In the last meeting Björn proposed to do correlations of cumulative increasing bins. He said to do that taking the zeroth point (last library point where prediction is still not done) and use it for having a potential 1 of correlation coefficient at the beginning. I could not do that because I didnt save the zeroth points, and this will be a bit tedious and confusing considering that many flies were tested and probably the order is not 100% known. Thus, I just did the bins skipping this zeroth point. After all, we should see something similar with this one. First two graphs: c105;;c232>TNT (first and second prediction point), second: WTBxTNT, third: WTBxc105;;c232.
Examples of how each of the flies look like. So they are basically cumulative bins with each single fly (each in different colour). Just to have a hint how does the singularity looks like. 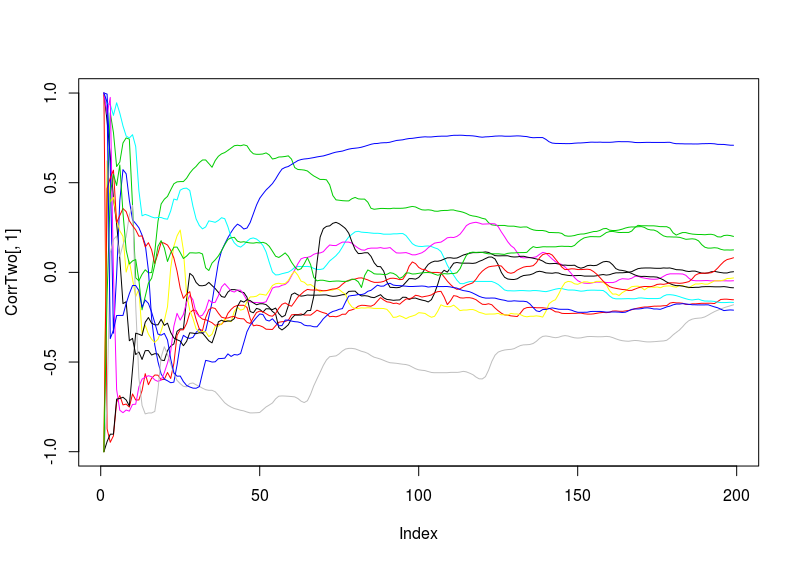
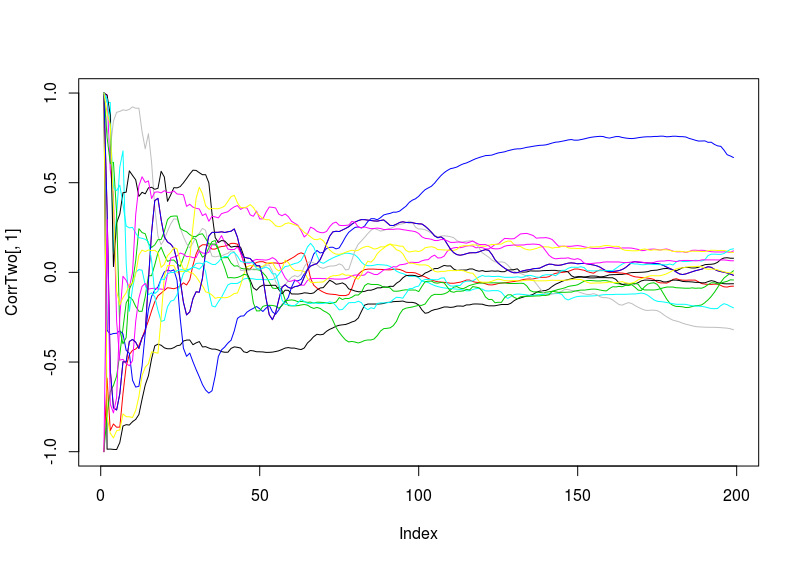
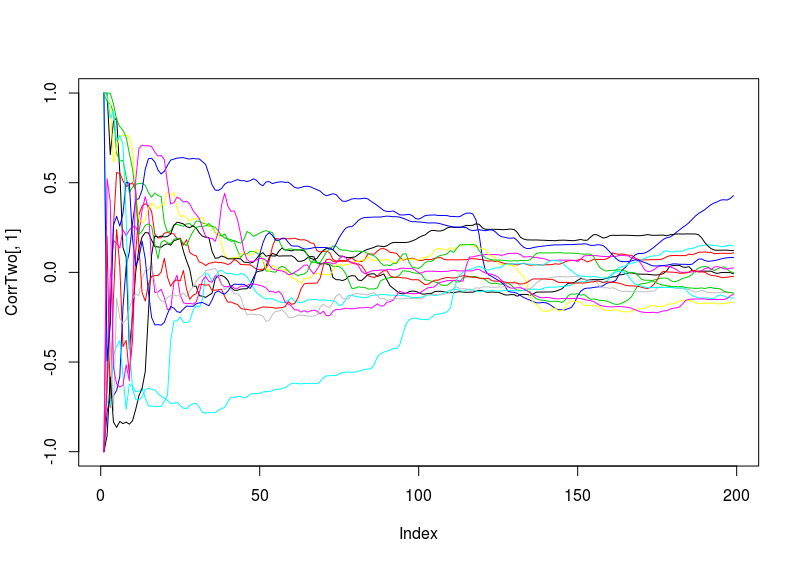 Second thing I did is normalize the to have a range from -1 to 1 all of them (I have to double check the range in the script) and also setting them at a starting point of zero. I did this because we do not want to have differences in the correlation coefficient due to a different offset of the values of the wing beat and neither because of the starting point (if the fly was already flying to the right full gas, then it could be that it has an influence in the following prediction).
Second thing I did is normalize the to have a range from -1 to 1 all of them (I have to double check the range in the script) and also setting them at a starting point of zero. I did this because we do not want to have differences in the correlation coefficient due to a different offset of the values of the wing beat and neither because of the starting point (if the fly was already flying to the right full gas, then it could be that it has an influence in the following prediction).
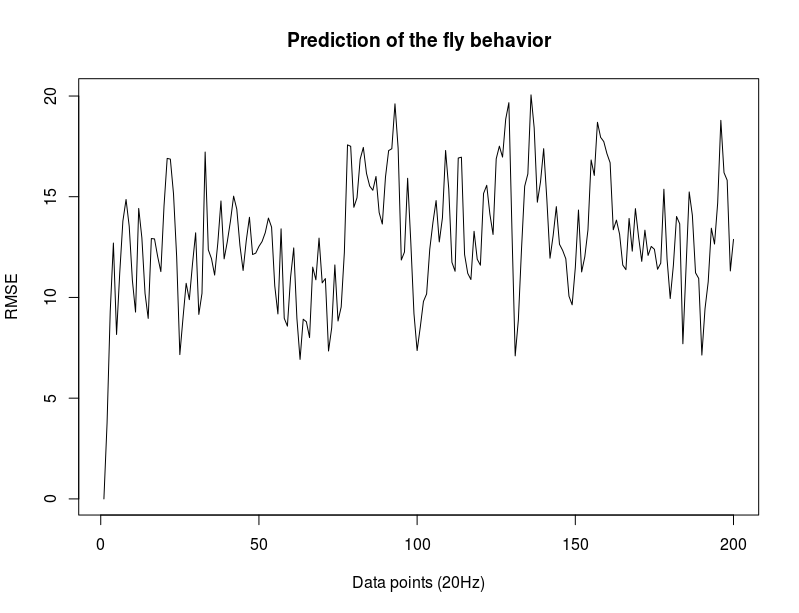
c105;;c232 –> first at starting at zero without normalizing and then with normalizing. The next is just the RMSE (not so important).
Category: R code, Spontaneous Behavior, strokelitude, WingStroke | No Comments
Prediction with binnning
on Monday, March 21st, 2016 1:13 | by Christian Rohrsen
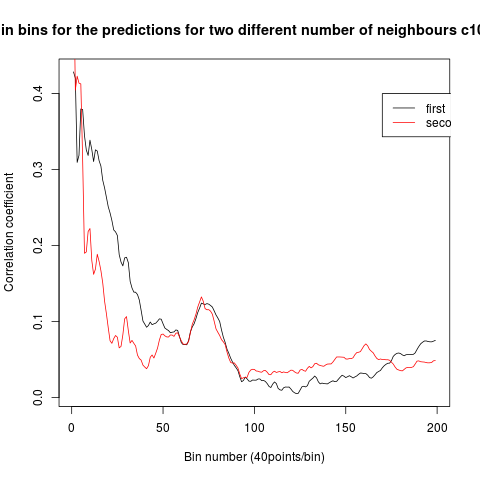
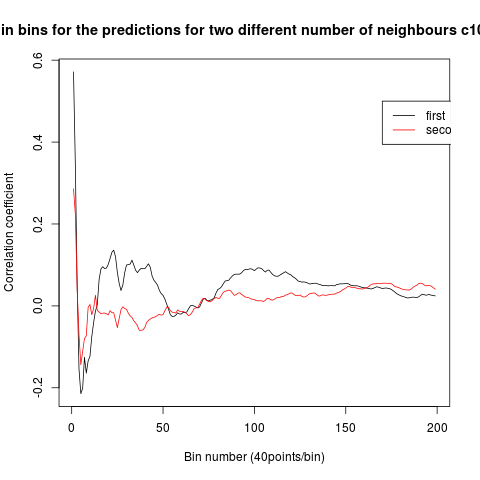
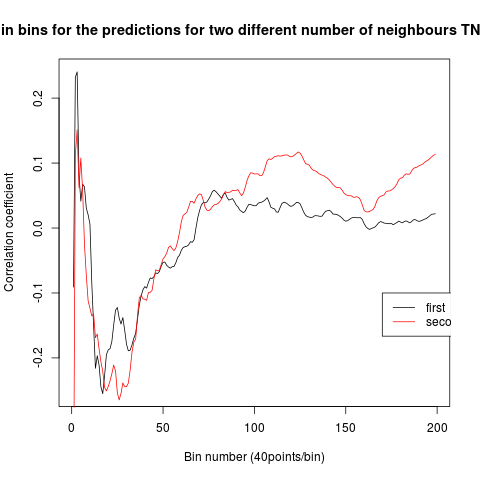
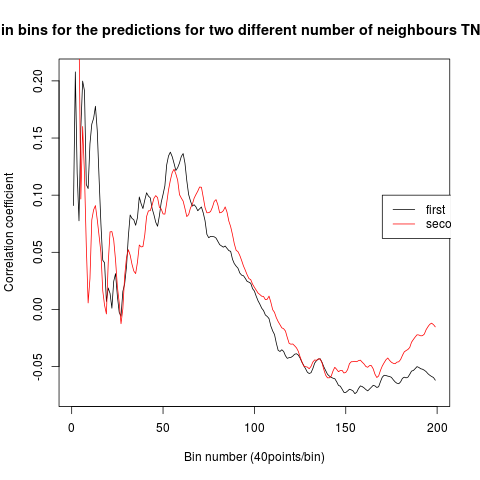
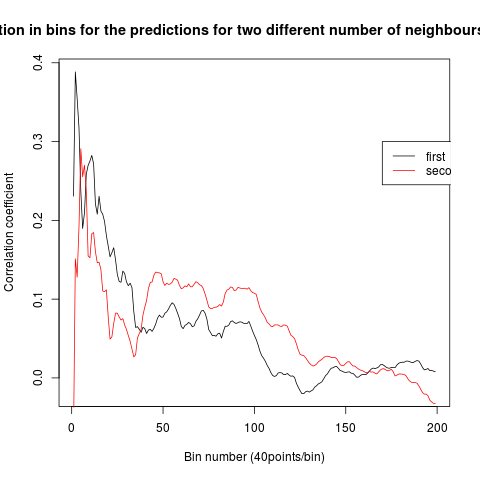
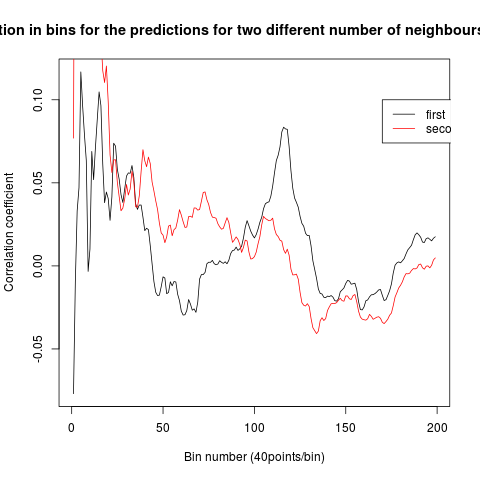
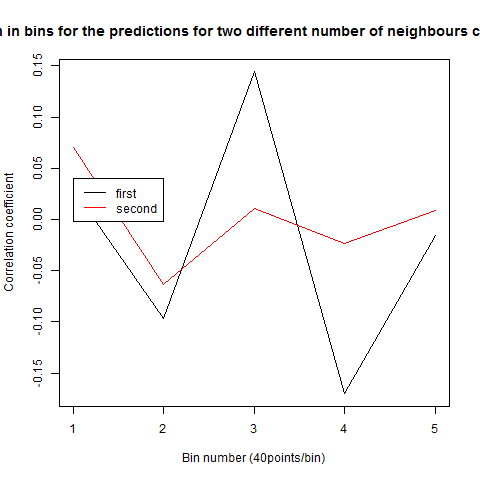
To see if there is an exponential decay in the prediction of the fly traces we did correlations of bins of 40 data points. We have 4 graphs (the last one merged) which consist of predictions at two different points with two different number of neighbours used for the prediction. So we have for each group sucesively: prediction at the first prediction point with the first number of neigbours, then the same with different number of neighbours. The last two are two different numbers of neighbours for the second prediction point. In the order: c105;;c232>TNT, TNTxWTB, c105;;c232xWTB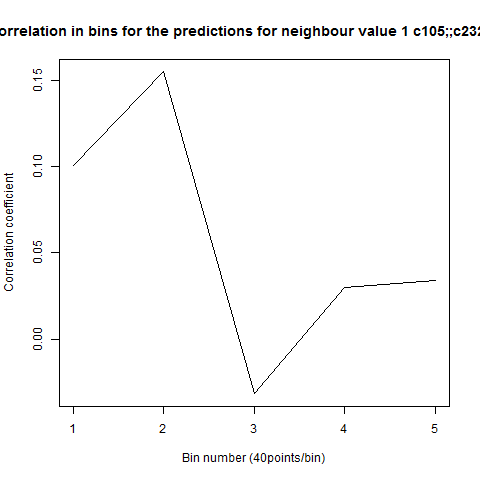
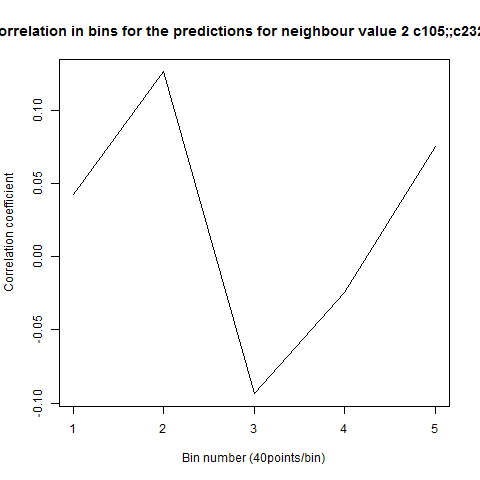
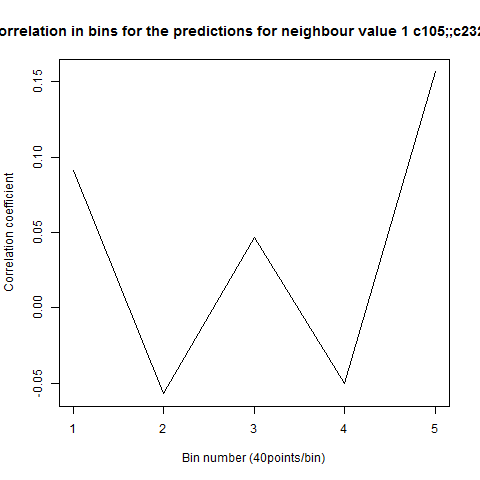
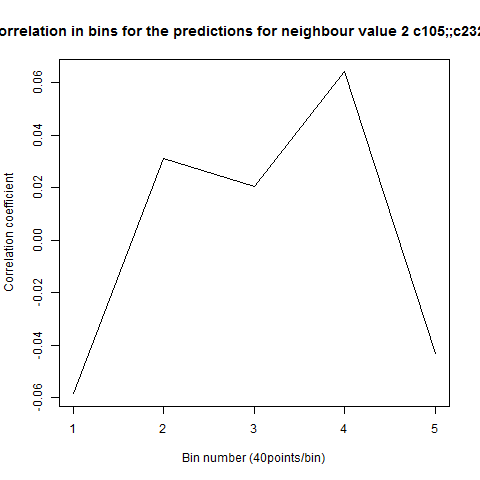
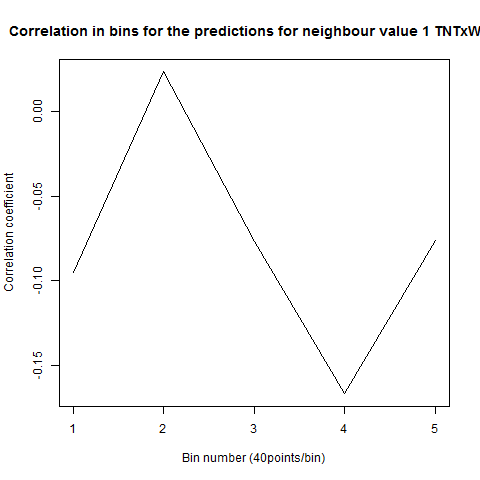
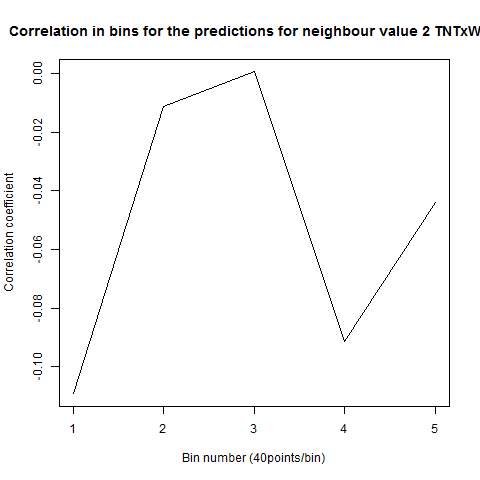
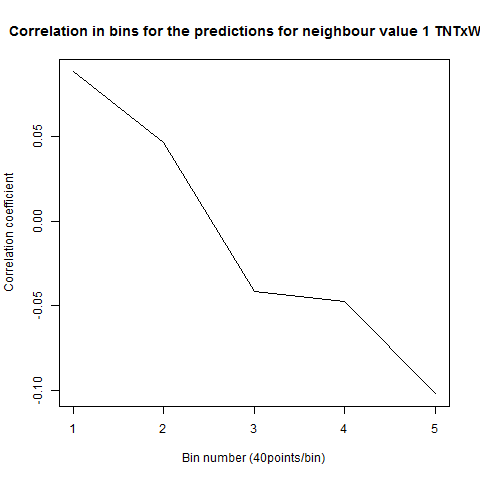
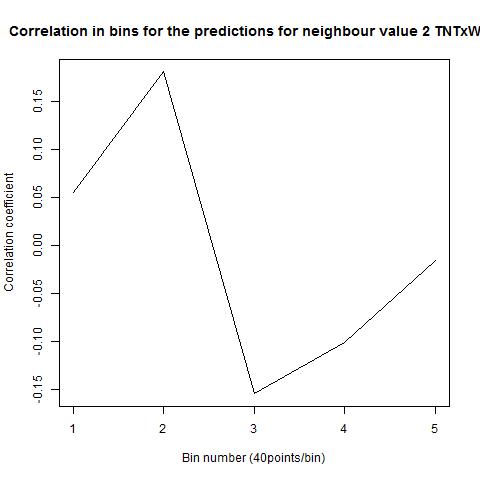
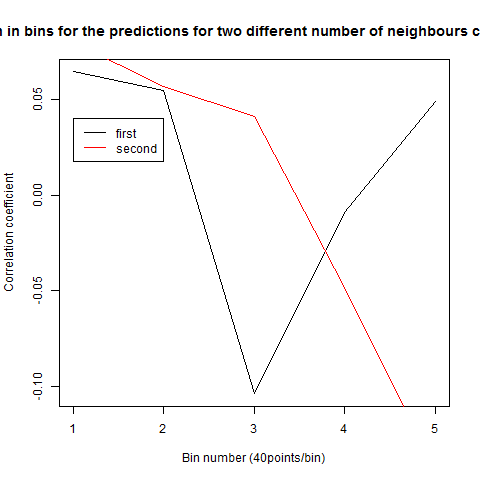
Category: R code, Spontaneous Behavior, strokelitude, WingStroke | No Comments